Hello
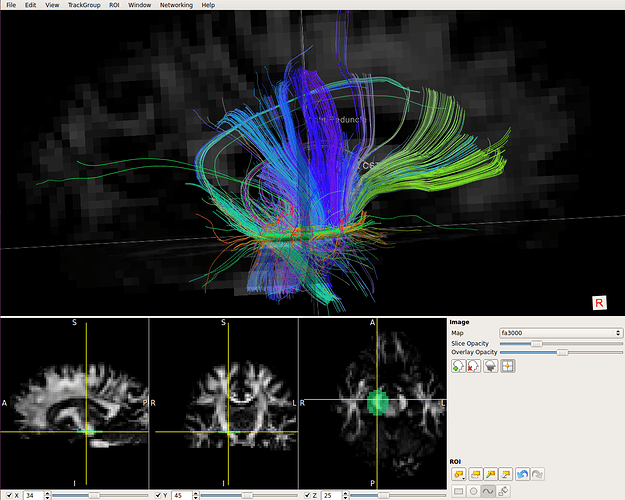
I am using CSD based deterministic streamline tractography to try and capture various structures (CST, IFOF, ILF, SLF etc), before applying ROIs to probabilistic streamlines. I am using standard approach, response from “fa” algorithm, b=3000 shell extracted from multishell dataset at the moment. Response, principal directions, FODs all quality checked etc, but when I do the deterministic streamline tractography, I am getting streamlines that truncate before I would expect:
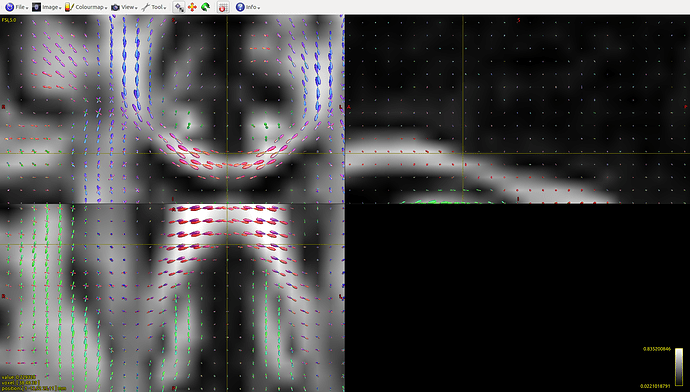
This is an example from a peduncle ROI, the streamlines seem to terminate at the level of the CC, the FODs there look like this:
I am using the command: tckgen -algorithm SD_stream -mask mask.mif -seed_image fathresh_0_2.mif -number 100000 streamlines.tck fod.mif
I’ve also tried different masks for the -mask option and different values for -cutoff option (0.01, 0.05) etc, this doesn’t seem to alter results.
Generally, beyond the CST, in many of the other fascicles, SLF, IFOF, UNC, etc, the streamlines seem to terminate generally fairly early, which makes them quite hard to capture using standard ROI dissection techniques, i.e. Catani 2002.
Does it look like I am doing something in error, or is this expected for MRtrix deterministic tractography?
Thanks in advance
Matt