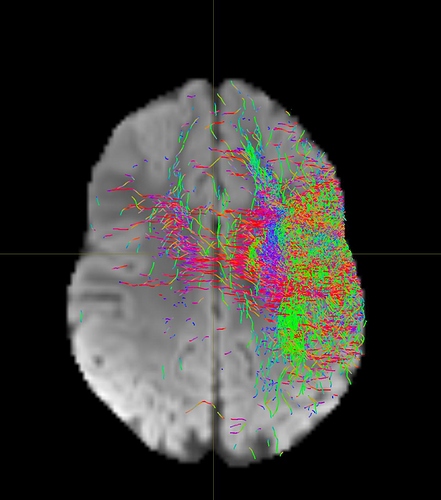
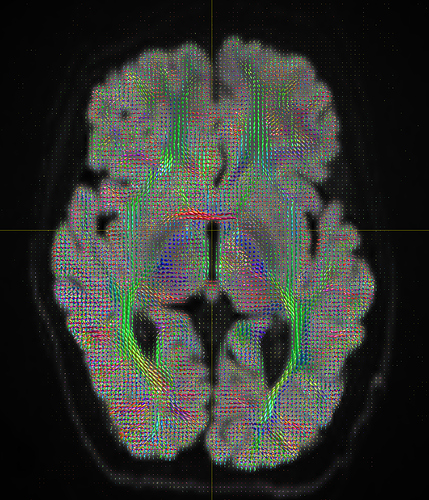
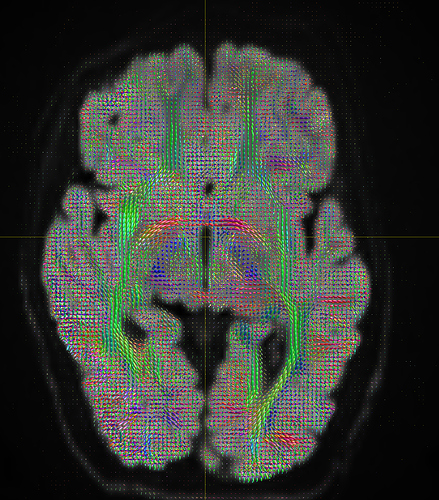
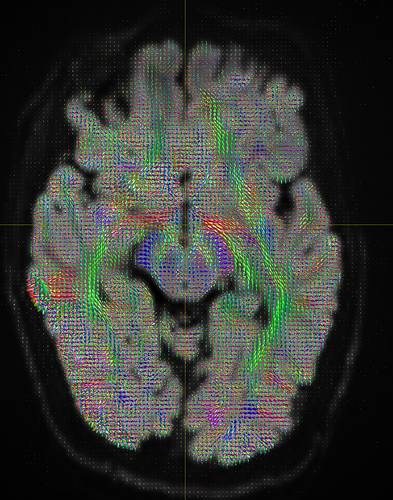
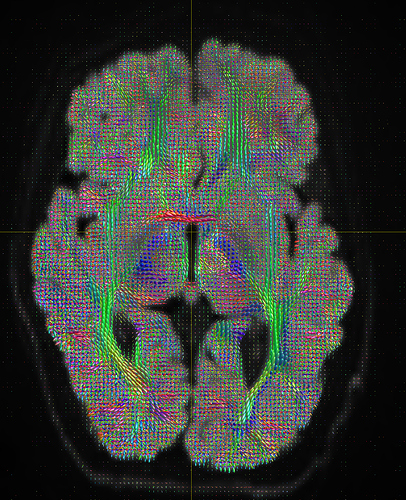
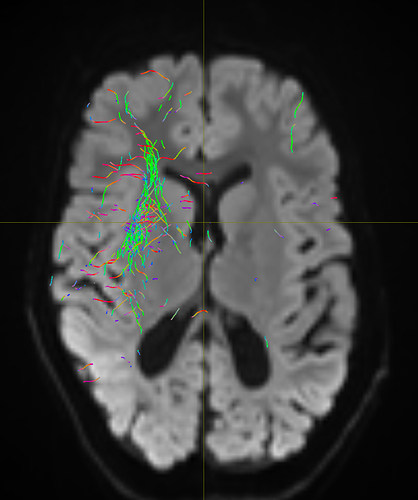
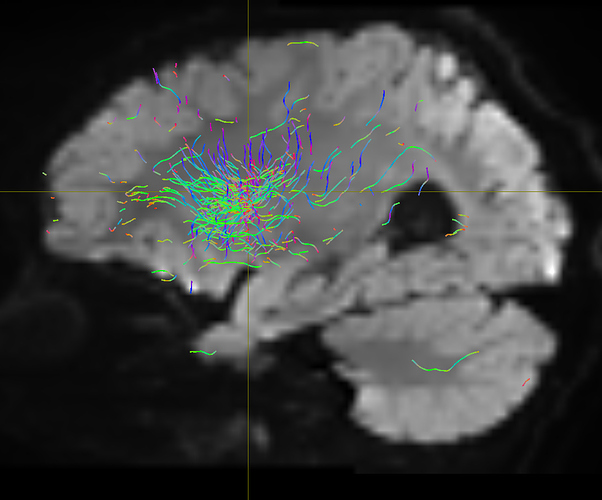
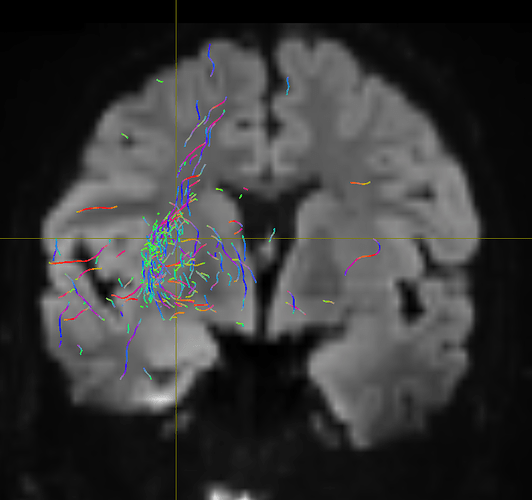
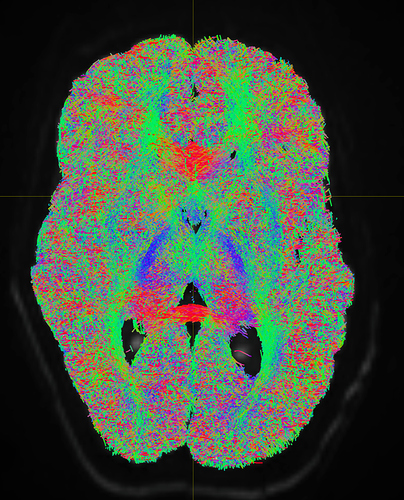
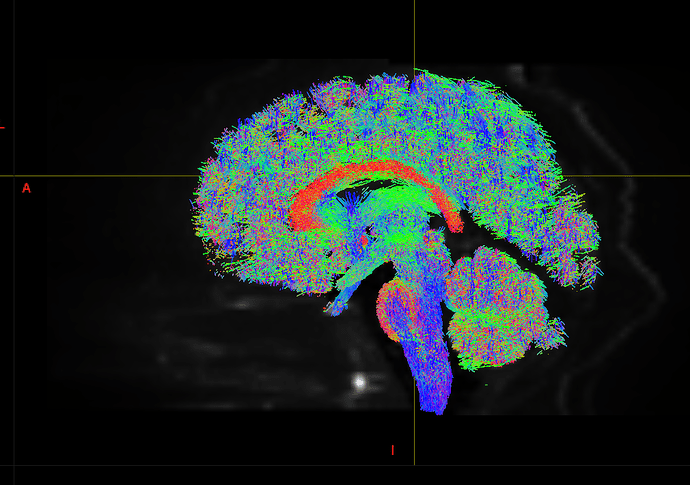
I am getting these types of fibres after running tckgen file, I don’t know what could be the possible problem but I think its in bvecs file. I generated bvecs file from the command export_grad command of mrtrix.
how to generate tracks as shown on mrtrix youtube tutorial.
Hi @nayan_wadhwani,
If you suspect an issue with your diffusion gradient table, try running the dwigradcheck script. It might provide some insight as to whether an erroneous axis permutation / flip has been introduced.
However:
I generated bvecs file from the command export_grad command of mrtrix.
If you are utilising MRtrix3 commands for tractography, there is no need to write the gradient table to a bvecs / bvals file pair, since the table will remain present in the image header; exporting those files is only necessary if you are invoking tools from some other software that requires those data in that format. There’s some more information on these data in the online documentation here.
Rob
My data is 64directions with and have 74images. I followed all the documentation steps that is given in the Mrtrix tutorial for the preprocessing and used -seed_position as a my roi while running an tckgen command. So my pinpoint question is what may be the mistake that I am making or what can be reason for these types of fibres. I followed tournier suggestion also by converting my data to 3D using mrconvert command but still there are same results.
Are you referring to this post? If so, it has nothing whatsoever to do with your issue here… My response was strictly to the original post, I’m not sure what could have given you the impression that it had anything to do with the issue you’re facing here?
If you’re trying to understand where your processing might have gone wrong or could be improved, I’d encourage you to post the exact series of commands that you used to process your data, right from the original DICOM import.
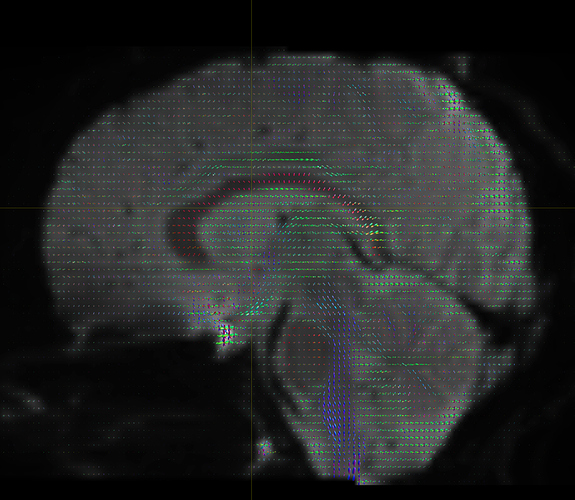
I’d also encourage you to look at your fibre ODFs in mrview’s ODF plot tool, and check that the orientations are as expected – if you’re in any doubt, please post a screenshot of the ODFs for us to have a look at.
Another thing to look at is whether your ROI is in the right place – this can go wrong depending on how it was generated. You can use mrview’s overlay tool to display it on top of the diffusion data (e.g. the FOD image as produced by dwi2fod) to verify that you are genuinely seeding from the place you were expecting.
But otherwise, I’m not sure what the problem is exactly. The results look noisy, but not outrageously so. You might find that using a higher -cutoff value in tckgen than the default will already clean up the results. But otherwise, this just looks like I’d expect probabilistic tracking to look like if seeded from a region like the centrum semiovale – lots of pathways going in lots of different directions in this region, it’s not surprising to find streamlines going all sorts of places. Of course, this all depends on what your seed region was exactly, it would help if you could post a screenshot of that too.
I am really Sorry I misunderstood the post. Okay Here I am posting all the list of all the commands as entered from top to bottom.
order of the command is bit different in the screenshots. here I am writing the correct sequence.
- dwi2denoise
2.dwimrdegibbs
3.dwibiascorrection - dwipreproc
- dwi2response dhollander gm.txt wm.txt csf.txt
- dwi2fod msmt_cst
- tckgen
Thanks Thijs for your valuable response,
and tried running tckgen command again, my command is
tckgen fod_wm.mif cst.tck -seed_sphere “position” -act 5TT.mif
after running its showing
500000 seeds 0 streamlines 0 selected
I used this command for generating my
I getting these types of fod’s
I haven’t tried ssmt-csd but i will definitly try once i solve this issue …
We are organising an neuroimaging workshop and in that we are doing hands on mrtrix session for dwi data.
That is why solving this issue is very important to me.
Ok, so that’s indeed 2-tissue MSMT-CSD then (if you need to describe at some point what it “is” you do here). This should at the very least deal with the initial problem.
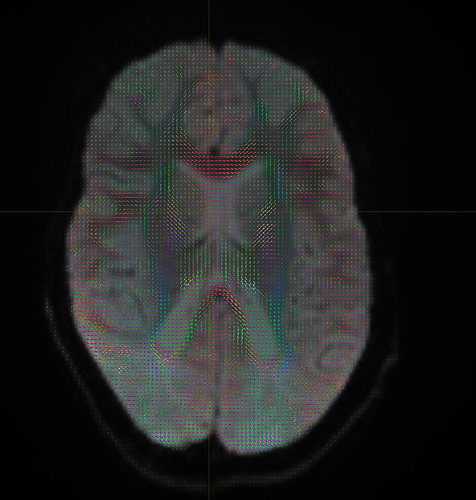
Yep, that checks out. Note that, compared to your earlier images, the CSF (e.g. ventricles) now has much lower intensity, and the FODs are either gone or drastically scaled down in those areas. That’s good, that’s definitely the intention of 2-tissue MSMT-CSD there. The FODs in the cortex and the intensity there are definitely still present, and they’re messy (as they would be at low b-value). However, that shouldn’t stop you from getting tractography results.
No worries, while it would help with the (cortical and other) GM, the main mistake you made before has been corrected.
So what remains is the question why you’re suddenly not getting streamlines. That’s gotta be either the amplitudes for some unexpected reason, or your seed sphere being in an impossible location, or the 5TT image not aligning (or otherwise itself being incorrect). Or else, something I’m missing of course.
So one step at a time: if you just run tckgen, seeding from the whole brain mask, and not specifying any seed spheres (just seed from the whole brain mask) and not enable ACT. If that doesn’t work, something’s wrong with the amplitudes. If it does work, something’s wrong with your seed sphere or maybe the 5TT volume.
Cheers,
Thijs
Hello Thijs,
I ran the whole process again using
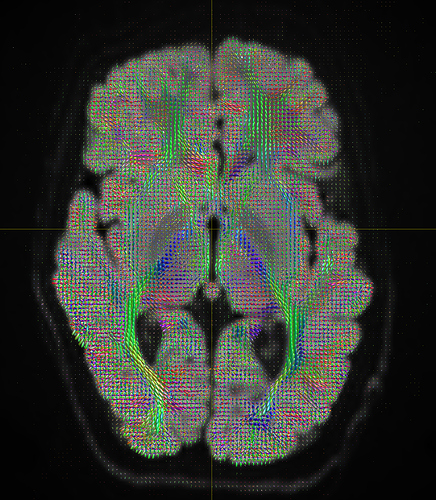
and generated tracks using whole brain mask here I am attaching the wholebrain tracks image and fod’s without using ACT and 5tt command and end results are still same.
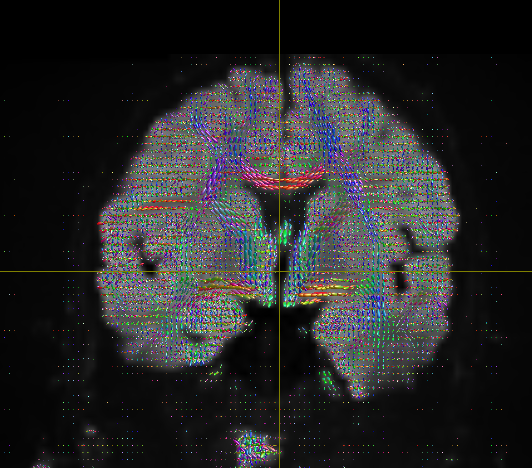
Here i have all the files that I generated using new commands that you have provided above but as you can see in the problem still persist. and I wanted to see tracks like this
Alright, so we’re at a point now where:
- the FODs are not broken anymore (before with the huge ones in the CSF, that definitely was not the case); improvement is likely with SS3T-CSD, but what you’re doing now at least isn’t technically wrong anymore (again, it was kinda “wrong” before, with those FODs in the CSF)
- your seed sphere or region seems fine: you get streamlines alright, and somewhere in that “mess”, you can spot traces of structure though; so you’re definitely seeding in a WM tract at least
- ACT seems to then be the thing/addition that suddenly makes all streamlines disappear, unless I missed something
- So the 5TT image might be wrong / problematic / misaligned / …
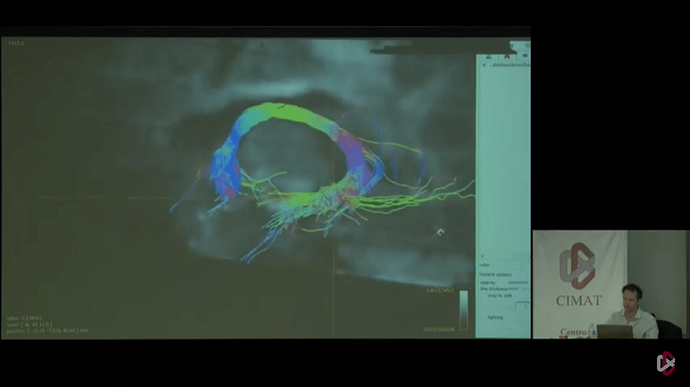
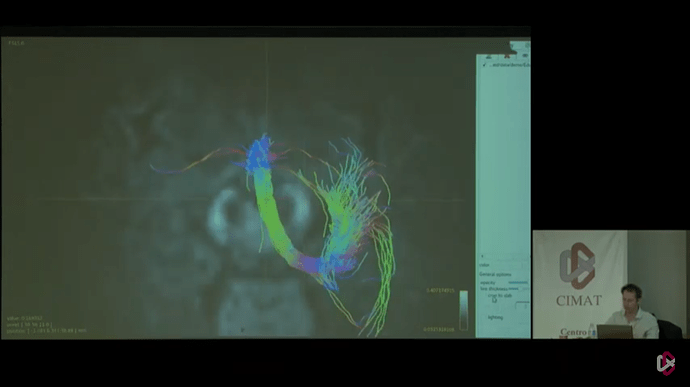
But in the end, I think the key to getting results like in those last 2 photos, lies in defining 2 reasonable (separate) “end” regions of the tract. Not really “end” regions per se, but just 2 way points that are quite a bit separated along the tract. That requires streamlines to at least trace the part of the tract between those regions, and makes it very hard for them to deviate from that path (because they will otherwise likely miss one or the other region). For an example like this, I wouldn’t add the complexity of ACT, to be honest. If you choose your “end” regions well, there isn’t much reason to add more complexity: you should be able to get a decent tract between the regions. Once you got those, you should add each “end” regions as a separate -include region to tckgen. That then requires each streamline to visit both regions to be accepted.
Seeing that it’s @jdtournier in those photos there, I reckon he might be able to help you or inform you about the regions he used to generate that particular tract on display there.
Thanks Thijs, For your convincing reply will run that with the tckgen command and will wait for the @jdtournier reply to generate those types of tracks.
Ok, I’ll see what I can add here…
First off, what b-value are you using? Lower b-values will make the results less impressive. Also, how old is the subject? Things get noticeably weirder with really young subjects.
As to what you can try, my first suggestion would be to increase the -cutoff in the tckgen call to something a bit higher than the default (e.g. 0.1 or even higher if it’s still messy). Tractography is a bit of a black art, there are many parameters that can be adjusted, and there is no right answer – it depends on your data and what you’re trying to do with it.
Also, can you post the full command-line you’re using in your tckgen call, just to check if there’s everything obvious?
Thanks tournier, Thankyou for your valuable response using the -cutoff value to 0.15 solved my problem and tracks are much better that it appeared to be before and I am really happy with the tracks that I am seeing.
Anyways my b=1000
tckgen fod_wm.mif -seed_sphere “position” file.tck -mask mask.mif -force
Good to hear. Although I don’t think we’ve solved the underlying issue really. You shouldn’t need such a high cutoff to get decent streamlines from data of this nature. But I can’t see anything too suspicious from the commands you’ve posted so far.
The only thing I can think of that might explain it would be if you’d estimated the response functions from the bias-corrected data, but applied them to the non-bias corrected data – that can introduce difference in scaling that might explain this. But from what you’ve posted, that’s not the issue…
I might also suggest that you run:
mtnormalise -mask mask.mif fod_wm.mif fod_wm_norm.mif csf.mif csf_norm.mif
And try using tckgen on the fod_wm_norm.mif file instead.