Dear all,
First of all, here’s an example of the code I’m using now to crop the fixels I give as input to the FACT tractography. Names of files are written in between <>. I now create a binary mask of fixels to keep (based on some voxel-wise relative amplitude threshold). I later apply this mask to crop the fixels (step 6).
% (1) Segment FOD to fixels (create fixelDir with a fixels-peak-amplitudes file)
fod2fixel -peak
% (2) Split the fixels to 3 volumes per fixel (fixelsNum x 3 file), later used for FACT tractography (create fixelPeakDirsFile file)
fixelFile = fullfile(fixelDir,‘directions.mif’);
fixel2voxel split_dir
% (3) Find the maximal amplitude of each voxel (create fixelPeaksMaxFile file)
fixelPeaksAmpsFile = fullfile(fixelDir,fixelPeaksAmpsFile);
fixel2voxel max
% (4) Map this maximal amplitude back to fixel space (fixelsNum x 1 file) (create fixelPeaksMaxFixelSpaceFile file)
voxel2fixel
% (5) Create a binary mask of the fixels, at 10% of the maximal FOD in each (create thresholdFixelsMaskFile file)
mrcalc <fullfile(fixelDir,fixelPeaksMaxFixelSpaceFile)> 0.1 -mult -gt 1 0 -if
% (6) Crop the fixels using the fixels mask (this creates a new fixelDir: croppedFixelDir)
fixelcrop
% (7) Extract the cropped fixels to a file with 3 volumes per fixel (fixelsNum x 3 file) (create croppedFixelPeakDirsFile file)
croppedFixelFile = fullfile(croppedFixelDir,‘directions.mif’);
croppedFixelPeakDirsFile = fullfile(croppedFixelDir,‘fixel_dirs3vols.mif’);
fixel2voxel split_dir
% (8) Run FACT tractography (create tckFile)
tckgen -algorithm FACT -seed_image -mask -select 500000 croppedFixelPeakDirsFile>
Similarly, I also create an absolute threshold mask of 0.2 amplitude.
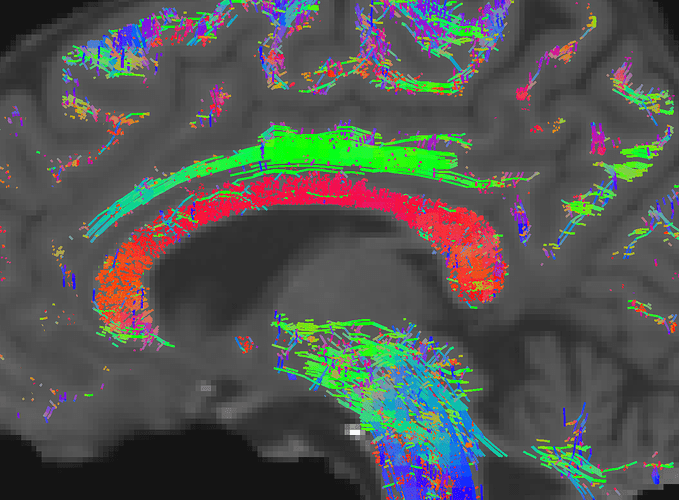
My problem now is that the tractography results seem to be contaminated by false positives that come from spurious fixels peaks. For example, in the corpus callosum many streamlines have anterior-posterior orientation (showing here dummy runs with a total of 50k streamlines):
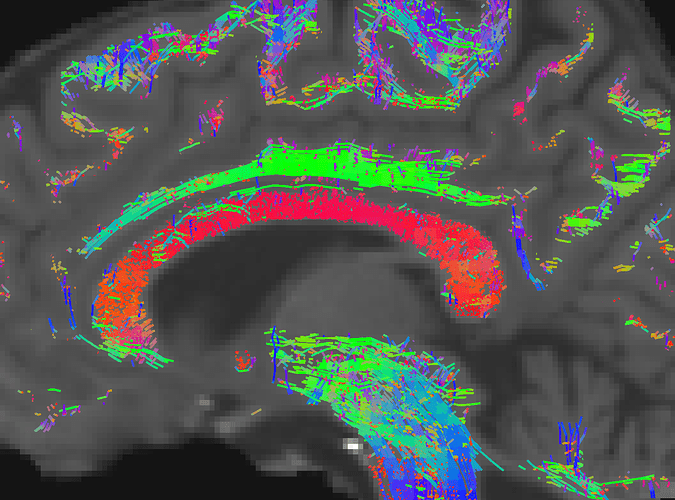
If I increase the cutoff in the tractography to 0.2 (instead of the default 0.05), I get similar results:
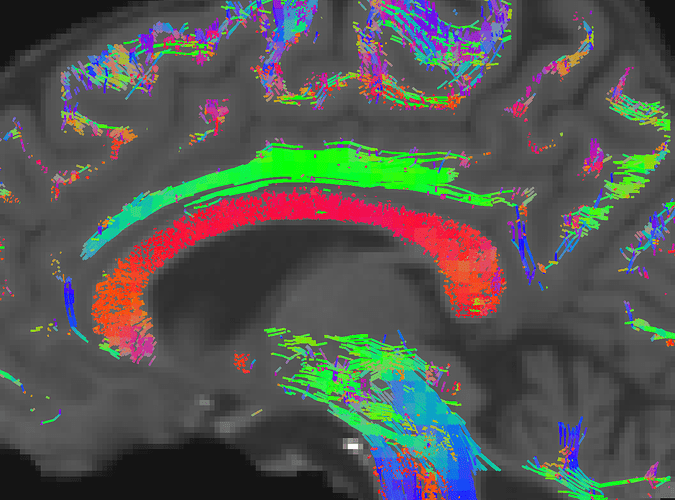
However, if I first crop the fixels using my absolute binary mask of 0.2 amplitude, the results are much cleaner:
Is this possibly related to the following post?
My question is: Do you have an idea why the cutoff option doesn’t do what I expect it to do?
In addition, judging by the cube-like structure shown in the tractography image above, I understand that perhaps I should have upsampled the diffusion FOD data before continuing to the next steps.
Finally, another short question. In many works (e.g., this one) I’ve seen that people also use an absolute threshold on the peak FOD, which is defined as 3 “multiples of the amplitude of a spherical FOD obtained from an isotropic voxel of gray matter”. Could you please tell me how you interpret this? My first thought was taking the l=0 volume of the FOD file, and average across gray-matter voxels (using the 5tt file, for example). However, this doesn’t seem right, and perhaps I need to convert this to an FOD amplitude somehow.
Sorry for the lengthy descriptions.
Many thanks!
Roey