Hello everyone,
I am not bringing up a new topic - but despite searching the existing entries, I did not find a solution for my problem. So I was hoping for some help for my analysis.
For using Tractography I recently started using MRtrix and tried to execute the 5ttgen command with fsl. This worked perfectly for many of my subjects using the T1weighted images. But for some subjects I got the following error code:
5ttgen: **[ERROR] FSL FIRST has failed; only 9 of 10 structures were segmented successfully (check /Data/DTI_test/sub-s22604/ses-1/dti/MRtrix/5ttgen-tmp-K6QCA2/first.logs)**
Checking the logs I get:
mrconvert '/Data/DTI_test/sub-s22604/ses-1/dti/MRtrix/T1_raw.mif' '/Data/DTI_test/sub-s22604/ses-1/dti/MRtrix/5ttgen-tmp-K6QCA2/input.mif'
mrconvert input.mif T1.nii -strides -1,+2,+3
maskfilter /fsl/data/standard/MNI152_T1_1mm_brain_mask_dil.nii.gz dilate mni_mask.nii -npass 4
standard_space_roi T1.nii T1_preBET.nii.gz -maskMASK mni_mask.nii -roiFOV
bet T1_preBET.nii.gz T1_BET.nii.gz -f 0.15 -R
fast T1_BET.nii.gz
run_first_all -m none -s L_Accu,R_Accu,L_Caud,R_Caud,L_Pall,R_Pall,L_Puta,R_Puta,L_Thal,R_Thal -i T1.nii -o first
Apparently “L_Accu” is missing. run_first_all was the last command where the analysis stopped.
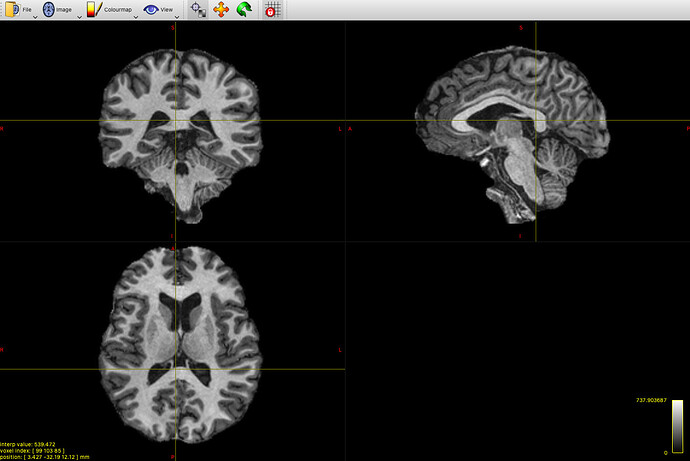
My input-image looks like this:
Does anyone have an idea what to do? How can I find out what is the problem?
Thank you so much in advance for all your help!
Best
Miriam