Hello experts
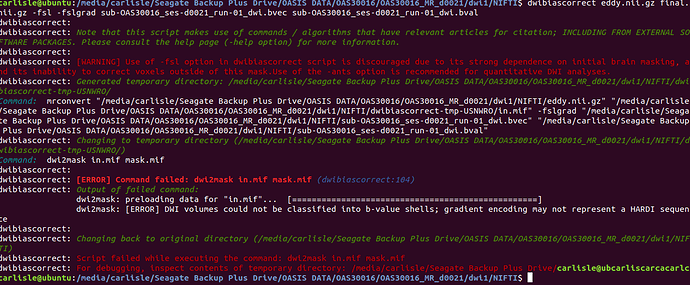
I’m doing dwi-bias correct… but When I use b-value 1000 direction 35 there is no error on it… but now I’m using multi b-value 0 50 350 600 900 1150 100 400 650…like this and direction is 25
and I use command option
‘dwibiascorrect input output -fsl -fslgrad file.bvec file.bval’
How can I solve it? Please help me…
It looks like the issue is that your data aren’t multi-shell HARDI. If I understand correctly, you have 25 directions spread over 9 b-values? If that’s right, the only thing you can do with this is DTI (using MRtrix3, at any rate). For dwipreproc, you need data that can be split into distinct shells each with a substantial number of directions – this is why that command is failing. But you’ll encounter similar issues at other stages in the processing. There’s not much you’ll be able to do with these data, I’m afraid…
Thank you for your answer!
Actually… I can’t clearly understand what is that split into distinct shells each with a substantial number of directions?
What is that mean?
Best regard, CS
In the context of high angular resolution diffusion imaging (HARDI), a ‘shell’ denotes the set of images (or volumes) that were acquired with the same b-value, but along different directions. To be any use, any (b>0) shell would need to have enough distinct directions to characterise the angular structure of the signal at that b-value. How many directions are required depends on b-value, amongst other factors. But these directions need to sample from the (half) sphere as uniformly as possible to avoid orientational bias.
Putting all that together, a multi-shell HARDI protocol might typically contain relatively few distinct b-values (maybe 3 or 4, e.g. b=0, 750, 1500, 3000 s/mm²), each with a decent number of directions, often with higher numbers for the higher shells (e.g. 5, 20, 50, 80 for the illustrative b-values above).
If I’ve understood your protocol correctly, there’s no way it can be described as a multi-shell HARDI sequence – but most of our analysis tools assume that’s how your data were acquired…