Hi, dear @jdtournier
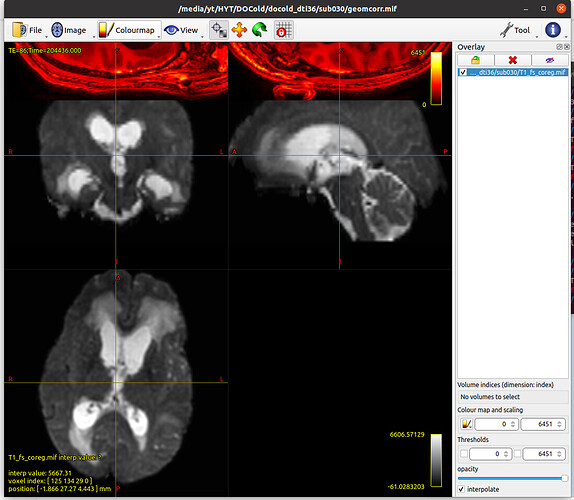
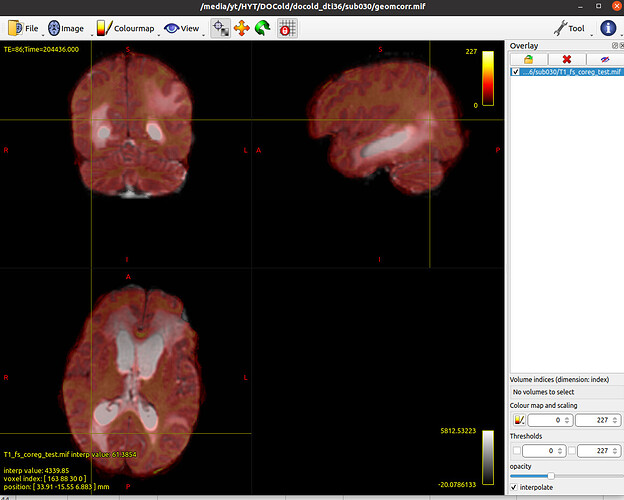
I’m trying to coregister the T1w to the diffusion image, but when I finished this and examine the quality of the coregister, something weird happened.
The coregistered T1w is far away from the DWI. Here are the commands:
# segmentation the T1w with freesuefer
recon-all -i *BRAVO*.nii.gz -subjid fs -all
# convert .mgz to .nii.gz
mrconvert fs/mri/aparc.a2009s+aseg.mgz aparc.a2009s+aseg.nii.gz
# 5ttgen with freesurfer algorithm
5ttgen freesurfer aparc.a2009s+aseg.nii.gz 5tt_fs_nocoreg.mif
(# I have tried the `5ttgen fsl` before, and the warnings showed with:
5ttgen: [WARNING] Generated image does not perfectly conform to 5TT format:
5ttgen: [WARNING] 5ttcheck: [WARNING] Image "result.mif" contains 8 brain voxels with non-unity sum of partial volume fractions
5ttgen: [WARNING] 5ttcheck: [WARNING] Input image does not perfectly conform to 5TT format, but may still be applicable (suggest re-running using the -masks option to see voxels where tissue fractions do not sum to 1.0)
, but when I switched to `5ttgen freesurfer` the warnings were gone.)
# convert .mif to .nii.gz
mrconvert 5tt_fs_nocoreg.mif 5tt_fs_nocoreg.nii.gz
# extract the first volume of the segmented dataset
fslroi 5tt_fs_nocoreg.nii.gz 5tt_fs_vol0.nii.gz 0 1
# coregister the two datasets
flirt.fsl -dof 6 -cost normmi -in mean_b0.nii.gz -ref 5tt_fs_vol0.nii.gz -interp nearestneighbour -omat dwi2t1_fs_fsl.txt
# transformation matrix
transformconvert dwi2t1_fs_fsl.txt mean_b0.nii.gz 5tt_fs_nocoreg.nii.gz flirt_import dwi2t1_fs_mrtrix.txt
# inverse the matrix to coregister the T1w to the DWI
mrtransform *BRAVO*.nii.gz -linear dwi2t1_fs_mrtrix.txt -inverse T1_fs_coreg.mif
(# Here I have tried both the skull-off and with skull T1w, and the effects are the same horrible)
Is there any problem with my commands? How can I solve this problem?
Thank you in advance!
Best,
Yuting.