Dear all,
I am combining two different atlases for connectivity analysis: Schaefer400_17networks and Melbourne Subcortical Atlas (S4). I manually created lut_in and lut_out by:
- Combining LUTs of both atlases (Schafer’s is 1001 to 2200, Melbourne’s is 1 to 54) for lut_in (_freesurfer.txt)
- Creating MRtrix version of lut_in with numbers starting from 1 to 454 for lut_out (_mrtrix.txt)
Those files can be accessible from this Cloud Storage.
However, running labelconvert results in some regions having high values (from “_freesurfer.txt”: e.g., 1093, 2094, 2087, etc.) in the magnitude of e+09. Of note, labelconvert runs without any warnings or errors.
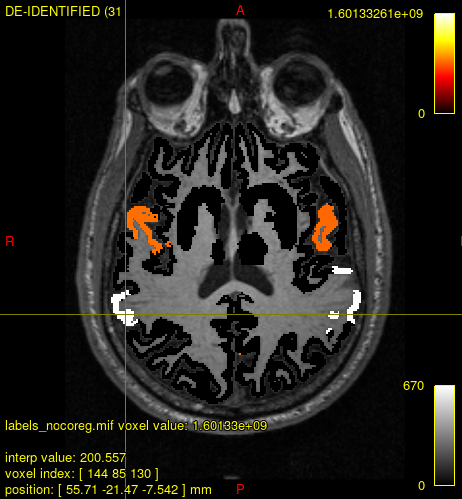
Any comments?
Best,
Amir