Hi Mrtrixers!
For my master thesis, I am performing the MSMT-FBA analysis exactly as described in Mrtrix-readthedocs, but
I noticed that my masks include a LOT of non-brain regions.
This is the case for both my binary, upsampled brain masks created for each subject (Step 6, dwi2mask), as well as the study-specific FOD template. (Step 9, population_template), the mask warped into template space (Step 11.1, mrtransform) and template mask (Step 11.2, mrmath). In addition, the binary template mask also contains some holes.
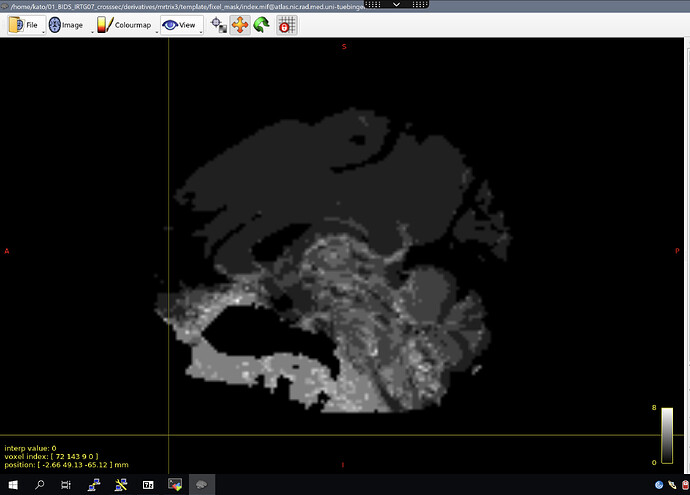
For my template analysis fixel mask (Step 12, fod2fixel), the problem is two fold; genuine whiter matter voxels seems to be excluded in the anterior and posterior part of the brain, but a large redundant part is also found at the bottom.
I think the problem starts in the first binary masks, and gets transmuted from their. How do I make the first upsampled brain masks fit better? How do I make the fixel mask fit better? Are additional steps needed to improve the other masks?
As for the upsampled brain mask images(step 6), it was mentioned that a generous mask was best, as regions that did not get included at this point, would not be used for FOD generation. However, for bias field correction and intensity normalisation, this same mask is used, but this time I am adviced to use a conservative mask (Step 9). How can these two pieces of advice be combined? Can I still use the same upsampled brain masks generated in step 6 for performing step 9?
Thanks a lot in advance!
Kato
Some additional information:
I preprocessed my data using a QSIprep container.
bval’s:
0 0 2500 1650 2500 850 1650 2500 1800 1650 850 1300 1650 2500 2500 1650 2500 850 0 1650 1850 2500 1700 850 850 1150 2500 2500 1650 2500 850 1650 1600 1600 1650 0 2500 850 1650 2500 2500 1650 2500 850 1650 2500 2500 1650 1600 850 1650 2500 0 2500 1650 2500 850 1650 1600 2500 1650 850 2500 1650 1600 2500 1650 1600 850 0 1650 2500 2500 1650 850 2500 1650 2500 2500 1650 850 2500 1650 1450 2500 1650 0 850 2500 1650 2500 2500 1650 850 2500 1650 2500 2500 1650 2500 850 1650 2500 0
mrinfo -size directions.mif:
864 964