Hi all ![]() !
!
I’m very grateful for this powerful platform. I have some questions when constructing a connectome using the AAL116 atlas. ![]()
-
Label Conversion
How should I correctly use labelconvert to map the original AAL116 labels (i.e., 2001-9170) to a compact 1-116 range?
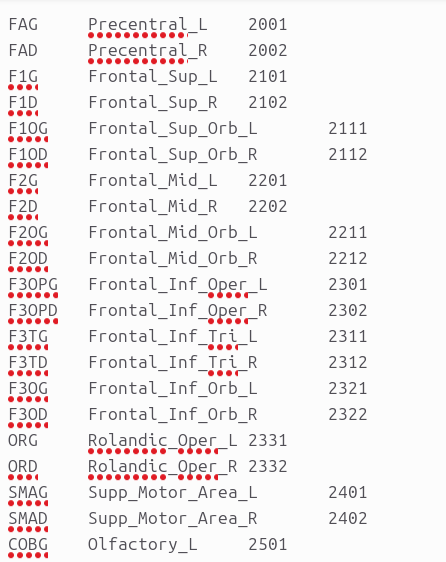
I used this: labelconvert AAL_in_DTI.mif aal116_lut.txt /usr/local/mrtrix3/share/mrtrix3/labelconvert/aal.txt AAL_in_DTI_relabelled.mif
Is my aal116_lut is correct?
-
Workflow Clarification
I’d like to confirm the best practice:
—1: First generate gmwmSeed_coreg.mif (from 5tt2gmwmi), then track streamlines, and finally apply the AAL116 atlas to construct the matrix.
—2: First create a seed mask directly from the AAL116 atlas (e.g., cortical ribbon + subcortical ROIs), then track and build the connectome.
Which order is recommended? -
Following Andy’s Brain Book’s pipeline step by step, my AAL116 connection matrix result is screenshotted below. Do these values look normal?
Any insights, references, or suggestions would be immensely helpful! Thank you in advance for your time and expertise.
Best regards,
Lane