Dear all,
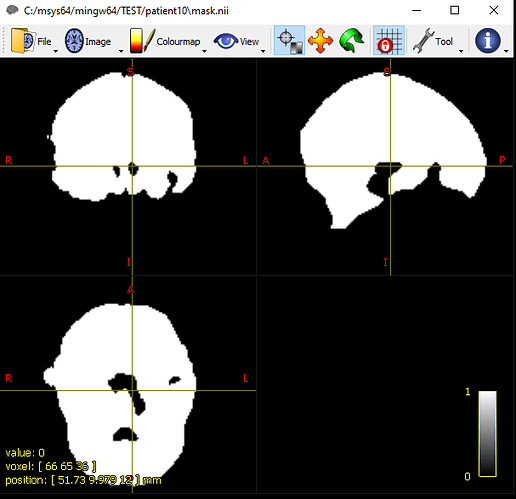
i have a probleme like a holes in the mask and i didn’t find the solution .
I really need your help and I would be happy for you.despite that I followed the link but I did not find the solution.
https://community.mrtrix.org/t/dwi2mask-holes-in-mask-images/484/16
Best Regards
Hi @zeraiiabderrazek,
Yes, this is not entirely uncommon. This tends to happen more in presence of challenging intensity inhomogeneities such as those caused by bias fields, but also very local ones due to EPI distortions. It’s even typically the latter that cause these holes to appear / be connected with the bottom of the brain, via a kind of “tunnel” that has an exit to the outside of the brain there.
Do look a bit around on the forum for what people commonly do, but generally pathways to a solution include:
-
Play around with bias field correction via dwibiascorrect and the parameters to the ANTs algorithm you can provide there. If you were already using dwibiascorrect here, try also without using dwibiascorrect as an alternative. It’s a complicated story (use the search function of the forum and look for dwibiascorrect for some other posts on this topic).
-
Originally, dwi2mask (as well as some later mask cleaning additions to that command) were designed for more typical and original dMRI spatial resolutions; e.g. 2.5mm isotropic or relatively close to that. At some point the FBA documentation was changed to apply dwi2mask to upsampled data, as this allowed for a slightly cleaner flow in the instructions; but since using that chance myself frequently, I’ve run into this issue with holes in the mask myself more often too. This might be a coincidence or subjective impression on my part, but in any case, I’ve changed my practice on this front recently back again. And maybe that was also a coincidence or subjective impression, but it appears I’m running less into the issue now as well. I’ve since been providing collaborators with updated pipelines that implement this change, and again I seem to be hearing about this issue far less now. Essentially, apply dwi2mask on your preprocessed but not upsampled dMRI data. If you require or desire upsampling in your pipeline, perform it afterwards, and regrid the mask at the original resolution equally to your upsampled resolution.
Other than that, this issue might still rear its head at times nonetheless. If that’s the case, assess how easy it would be to fix manually or apply well engineered series of commands to apply e.g. morphological operations to your masks to get this resolved in most (if it affects many masks in your study). If all that still fails or is too cumbersome, check whether you’ve got other ways of obtaining reasonable brain masks, e.g. from a T1w image and using FSL’s BET software. In the latter case, you’ll probably also have to resample the resulting mask to the working resolution of the dMRI data in your study.
So well, no one size fits all solutions here… this requires a bit of tinkering at times, and the best solution might depend on your particular data and its qualities at hand…
Hope that helps!
Cheers,
Thijs
dear @ThijsDhollander
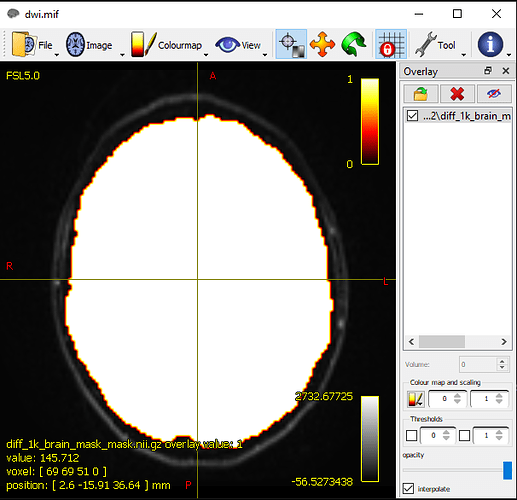
Thank you very much for your quick response,here I tried to do everything but the one she will work very well with my data DWI is by using FSL’s BET software.so you will find below the result of generation of the mask,I think that is the best solution because the use of the command dwibiascorrect actually depends on a mask so it’s closed loop.

so I hope this result is the best.
I’m waiting for your judgment:smiley:

Best Regards
1 Like
I can definitely confirm that’s a good looking mask there.  If that works well across all data you’re looking at, I’d advise to stick with that approach by all means!
If that works well across all data you’re looking at, I’d advise to stick with that approach by all means! 
Correctly identified, and indeed: bias field correction and mask estimation are in many ways a joint thing, with a chicken-and-egg problem element to it. However, bias field estimation can often get by with a sub-optimal mask. Especially the properties of that initial mask with the holes aren’t per se dramatic for that purpose: the parts of the mask that were present kind of “envelop” the hole that’s missing. Since a bias field is smooth, it would essentially be interpolated within that hole, based on data “around” it. So that’s why sometimes, it can help to perform bias field correction (with a sub-optimal mask) in order to then estimate a decent mask. You could even then use that better mask to re-estimate the bias field; and even keep on iterating both to “jointly” estimate them.
…however, in any case, you’ve got a good strategy now that works regardless; so I’d go with that. 
Cheers,
Thijs
1 Like