Hello everyone =) I would like your thoughts and suggestion on an issue that has been eating my brain lately regarding my scans.
I have been working with ex vivo animal scans (minimal movement, minimal signal loss) and under the supervision of an experienced external colleague. We have acquired many DWI scans with the aim of performing some basic (analyses like extracting FA, MD, RD values), and hopefully some advanced ones in the future (Fixel-based analyses). I am mostly new to MRI analyses so this has been a bumpy road plagued with uncertainties and a learning curve far from over.
My colleague advised to me about the general pipeline we would use to pre-process the data, which is roughly:
- Convert raw data to obtain nifti, bval and bvec files. I used
nanconvert_brukerfor this. - Transform niftis to mif files to integrate the gradient with the scans. I used
mrconvertfor this. - Perform necessary rotations and transformations using
mrconvert -axesandmrtransform -flipwhen necessary - Perform denoising and removing Gibbs ringing artifacts. I used
dwidenoiseandmrdegibbsfor these. - Prepare the necessary files and perform
eddy. I did this usingeddyrather thandwifslpreprocsince it’s what was available for me in the HPC. - Perform DTI analysis. I used
dtifitfor this.
I am currently sitting between steps 5 and 6 as I await for further instructions from my CI regarding what will our ROIs be. I asked my supervisor about if we’d need to use topup for our scans, since it’s a tool I frequently see other people using in their DWI pre-processing pipeline, but she said that we wouldn’t need it so we could skip it. So I did. The brains don’t look distorted as they do in the topup UsersGuide so I guess I can see why she said we wouldn’t need it.
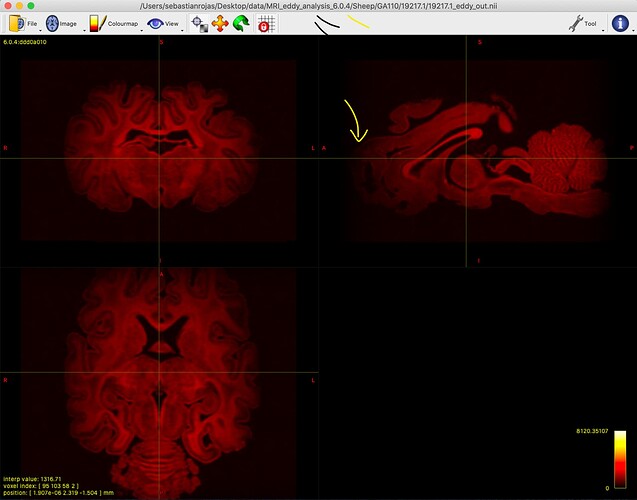
When looking at my data, I see certain parts of the brain that are “dimmer” than others (see below). These are usually in the outer parts of the brain. In this case, it’s in the anterior part of the brain. Frequently, it’s in the ventral part, and sometimes in other parts like the posterior side. Sometimes it’s in more places than just one. But it’s always at the outer edges of the brain, especially if the brain is larger (like in this case for a late gestation sheep) compared to a mouse.
In all my ignorance, I am pretty sure these inconsistencies need correcting before calculating diffusion parameters. but in all honesty I simply don’t know.
Can I ask you: What are these inhomogeneities called? Do you think these need correction as part of my DWI analysis pipeline? If so, what tool would you recommend to correct them, and at what step of the pipeline should I need to perform these corrections?
Also feel free to comment on the pipeline in general if you think I’m missing something.
Thanks for your constant help =) I really appreciate it.
Cheers,
Seb.