Hi MRtrix Users/Devs ! ![]()
![]()
Sorry to bother everyone like this but I’ve stumbled upon some issue with the following line of “6.Connectome Visualization tool” from BATMAN Tutorial :
<mrview hcpmmp1_parcels_coreg.mif –connectome.init
hcpmmp1_parcels_coreg.mif –connectome.load hcpmmp1.csv>
It is looping on “radeon : mmap failed, errno 12”
Then followed by “mrview: [SYSTEM FATAL CODE: SIGSEGV (11)] Segmentation fault: Invalid memory access”
I tried already running the command with gdb to see step-by-step where it crashed and I obtained the following results :
-
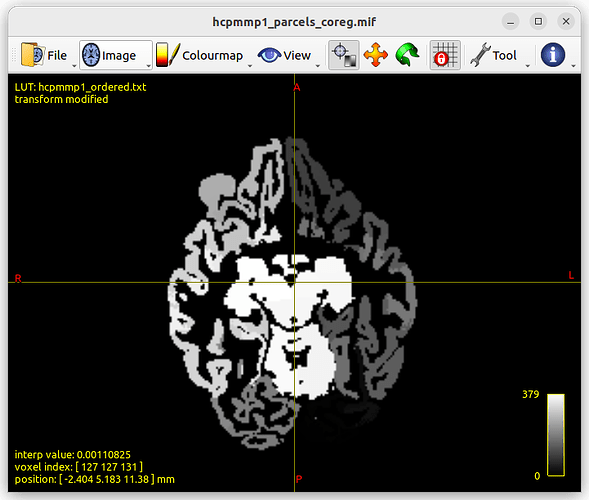
With only <gdb --args mrview hcpmmp1_parcels_coreg.mif>, I get to see hcpmmp1_parcels_coreg.mif in mrview (brain slices with different gray values depending on the areas ?) like this
-
With <gdb --args mrview hcpmmp1_parcels_coreg.mif -connectome.init \ hcpmmp1_parcels_coreg.mif> I get a black window and then the failure as aforementioned.
-
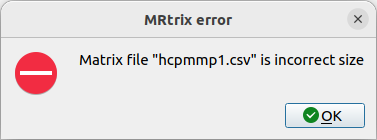
With <gdb --args mrview hcpmmp1_parcels_coreg.mif -connectome.load \ hcpmmp1.csv> I got this : (same results with my own csv or after using the one from Supplementary Files…)
& The same window as shown previously in (1).
Otherwise I managed to do the rest of the tutorial without too much issues (had one with mrview where I was missing GLIBCXX_3.4.30).
I am currently working on Linux (Ubuntu 22).
If anyone knows “the trick” to solve this it would help me a lot in my endeavours !
A.ROSSEZ