Hi there,
I am using the following code to
- map the annotation files of the Glasser atlas from fsaverage to the subject
- map the Glasser annotations onto the volumetric image and add (Freesurfer-specific) subcortical segmentation
- Replace the random integers of the Glasser file with integers that start at 1 and increase by 1
- register the ordered atlas-based volumetric parcellation to diffusion space
- generate connectome
#map the annotation files of the HCP MM1.0 atlas from fsaverage to the subject (first left, then right hemisphere)
mri_surf2surf --srcsubject fsaverage --trgsubject $subjectID --hemi lh --sval-annot $SUBJECTS_DIR/fsaverage/label/lh.hcpmmp1.annot --tval $subjectDIR/label/lh.hcpmmp1.annot
mri_surf2surf --srcsubject fsaverage --trgsubject $subjectID --hemi rh --sval-annot $SUBJECTS_DIR/fsaverage/label/rh.hcpmmp1.annot --tval $subjectDIR/label/rh.hcpmmp1.annot
#Map the HCP MMP 1.0 annotations onto the volumetric image and add (Freesurfer-specific) subcortical segmentation.
#Convert the resulting file to .mif format (use datatype uint32 --> liked best by mrtrix)
mri_aparc2aseg --old-ribbon --s $subjectID --annot hcpmmp1 --o $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1.mgz"
mrconvert --datatype uint32 $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1.mgz" $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1.mif"
#Replace the random integers of the hcpmmp1.mif file with integers that start at 1 and increase by 1
labelconvert $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1.mif" /home/julia/mrtrix3/share/mrtrix3/labelconvert/hcpmmp1_original.txt /home/julia/mrtrix3/share/mrtrix3/labelconvert/hcpmmp1_ordered.txt $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1_parcels_nocoreg.mif"
Register the ordered atlas-based volumetric parcellation to diffusion space
first calculate transformation
flirt -in $derivatives_path"/dwi_oct2020/"$sub"/"$ses"/dwi/"$subjectID"_dir-APPA_meanB0brain.nii.gz" -ref $derivatives_path"/dwi_oct2020/"$sub"/"$ses"/anat/"$subjectID"_acq-mprage_T1wBrain.nii.gz" -dof 6 -omat $derivatives_path"/dwi_oct2020/"$sub"/"$ses"/dwi/"$subjectID"_diff2struct_fsl.mat"
transformconvert -force $derivatives_path"/dwi_oct2020/"$sub"/"$ses"/dwi/"$subjectID"_diff2struct_fsl.mat" $derivatives_path"/dwi_oct2020/"$sub"/"$ses"/dwi/"$subjectID"_dir-APPA_meanB0brain.nii.gz" $derivatives_path"/dwi_oct2020/"$sub"/"$ses"/anat/"$subjectID"_acq-mprage_T1wBrain.nii.gz" flirt_import $derivatives_path"/dwi_oct2020/"$sub"/"$ses"/dwi/"$subjectID"_diff2struct_mrtrix.txt"
mrtransform -force $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1_parcels_nocoreg.mif" -linear $derivatives_path"/dwi_oct2020/"$sub"/"$ses"/dwi/"$subjectID"_diff2struct_mrtrix.txt" -inverse -datatype uint32 $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1_parcels_coreg.mif"
mrstats $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1_parcels_nocoreg.mif"
mrstats $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1_parcels_coreg.mif"
#connectomics
connectome_filename=$derivatives_path"/dwi/"$sub"/"$ses"/dwi/"$subjectID"_connectome_Glasser.csv"
tck2connectome $derivatives_path"/dwi/"$sub"/"$ses"/dwi/"$subjectID"_iFOD2.tck" $derivatives_path"/dwi/"$sub"/"$ses"/anat/"$subjectID"_hcpmmp1_parcels_coreg.mif" $connectome_filename --assignment_radial_search 3.2 -zero_diagonal --force
For some sessions, I am getting the following warning (with changing node numbers):
tck2connectome: [WARNING] The following nodes are missing from the parcellation image:
tck2connectome: [WARNING] 120
tck2connectome: [WARNING] (This may indicate poor parcellation image preparation, use of incorrect or incomplete LUT file(s) in labelconvert, or very poor registration)
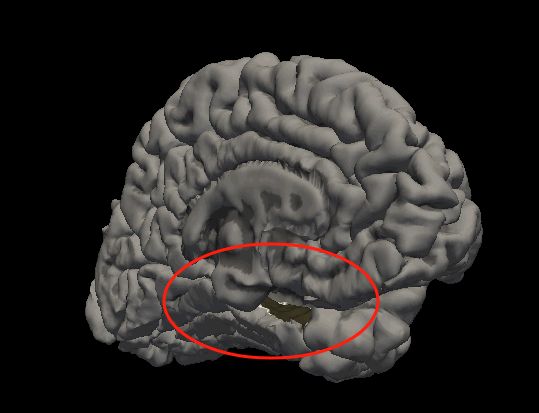
I checked the parcellation and it’s fine. What could be causing this?
thanks in advance for any help!
Julia