Hello!
I received a DKI dataset, consisting of two DICOM folders: one with only 3 volumes (b0s) and another with 102 volumes (with 12 b0s distributed throughout the volumes), bvec and bval. However, despite the indication of AP and PA, respectively, in the file names, when I view the files, I get opposite information.
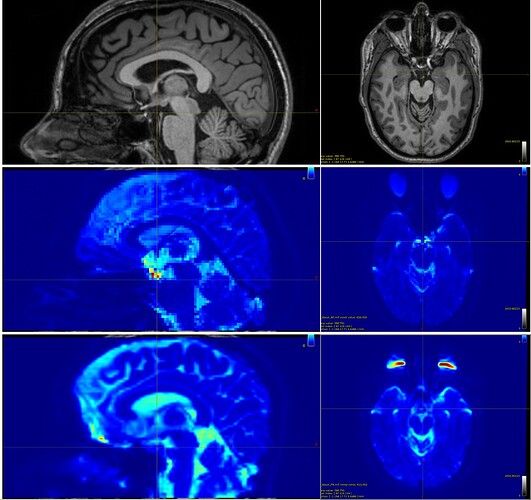
As you can see in the figure, the first row contains, for the same patient, the sagittal and axial T1-weighted images, followed by the same images (row two) for one of the DICOM folders (in this case, the one with 3 b0 volumes, supposedly AP) and, finally, by the same images (row three) for the other DICOM folder (with 102 volumes, bvec, bval and, supposedly, PA).
However, when checking the images, we see that, for the second row, the eyes are elongated forward, the nose is compressed, the scalp in the occipital region is expanded, and the cerebellum is slightly elongated, which actually corresponds to a PA acquisition. In the third line, we see the opposite patterns: the eyes flattened towards the brain, the nose expanded forward, the cerebellum compressed, and the scalp in the occipital region “thinner,” which would correspond to AP (contrary to what the name saved in the file suggests).
These characteristics were observed in the acquisitions of other patients. Therefore, I believe the names AP and PA were switched in the description. What do you think? Could you guide me?
Additionally, considering that the switch actually happened and the main file with the 102 volumes the PE = AP. I applied the following commands:
mrconvert sub01_AP/ sub01_AP.mif
mrconvert sub01_PA/ sub01_PA.mif
dwidenoise sub01_AP.mif sub01_AP_den.mif
dwidenoise sub01_PA.mif sub01_PA_den.mif
mrdegibbs sub01_AP_den.mif sub01_AP_preproc_input.mif
mrdegibbs sub01_PA_den.mif sub01_PA_preproc_input.mif
dwiextract sub01_AP_preproc_input.mif - -bzero | mrmath - mean mean_b0_AP.mif -axis 3
dwiextract sub01_PA_preproc_input.mif - -bzero | mrmath - mean mean_b0_PA.mif -axis 3
mrcat mean_b0_AP.mif mean_b0_PA.mif -axis 3 b0_pair.mif
dwifslpreproc sub01_AP_preproc_input.mif sub01_dwi.mif -pe_dir AP -rpe_pair -se_epi b0_pair.mif -align_seepi -eddy_options " --slm=linear "
dwibiascorrect sub01_dwi.mif sub01_dwi_unbiased.mif -bias bias.mif
I would like to know if the approach is correct (extracting the b0s and calculating the average for each DICOM folder, followed by concatenating the average b0 of AP and the average b0 of PA).
When I ran the dwifslpreproc command, two warnings appeared that I would like to know how problematic they are and if there is anything I can do:
dwifslpreproc: [WARNING] It appears that the spin-echo EPI images used for inhomogeneity field estimation have a different repetition time to the DWIs being corrected. This may cause issues in estimation of the field, as the first DWI b=0 volume will be added to the input series to topup due to use of the -align_seepi option.
dwifslpreproc: Balanced phase-encoding scheme detected in SE-EPI series; volume 0 will be removed and replaced with first b=0 from DWIs
Finally, I also have a problem due to cropping in the cerebellar region during acquisition. I’m thinking of using a standard mask in the MNI space without the cerebellum for group analysis with FA maps, for example. However, when normalizing the post-processing images to the MNI space and then using the mask without the cerebellum, could cropping in the cerebellar region be problematic? Any suggestions on how to proceed?
Kind regards!
 ) correct me…
) correct me…