Hi MRtrix crew,
thanks for developing and maintaining the tool.
I am trying to perform probabilistic tractography on an in-house high-res (0.7 mm iso; b=1000 s/mm2; 60 directions plus 1 b0) dMRI dataset. I am using iFOD2 through a Singularity container built from the 3.0.8 Docker container.
I am interested in the cerebellar region, but am having trouble getting a result that satisfies me. My ODFs look OK, but when visualizing the tracking results, I get mostly broken streamlines, and my SCP and MCP look extremely poor. I am selecting 3.5 M streamlines in a bounding box that encloses my regions of interest.
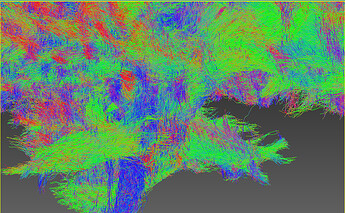
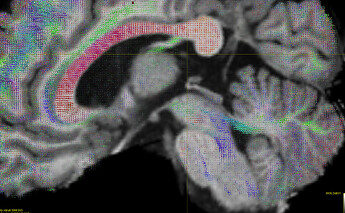
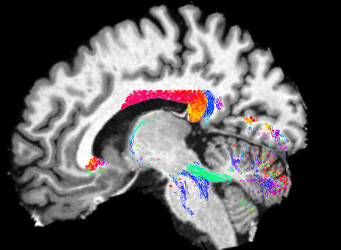
FODs:
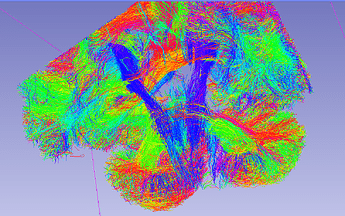
The below screenshot shows a subset (1%) of the 3.5 M streamlines:
Although it is a small fraction it shows the trend IMO.
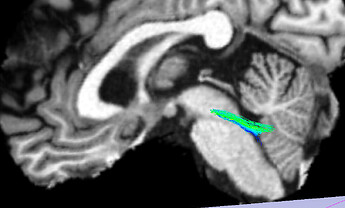
The extracted MCP looks like this (the sagittal cut is at the midline):
So not quite right either, as streamlines climbs through the SCP and do not travel across the pons. My extraction should not be a problem since it is working OK on other tractography data.
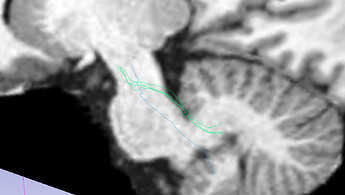
SCP:
My call looks like the following:
singularity run \
-B ${host_path}:${container_path} \
${mrtrix_apptainer_fname} \
tckgen "${in_fname}" "${out_fname}" \
-algorithm iFOD2 \
-select ${seed_count} \
-step ${step_size} \
-minlength ${min_length} \
-maxlength ${max_length} \
-seed_image ${seeding_mask} \
-mask ${seeding_mask}"
with
min_length=6
max_length=340
seed_count=3500000
step_size=0.2
Changing the step size (I have gone down to 0.01; screenshot above) does not have the impact that I would hope for a large change.
I know my min length is small, but there are streamlines in the region I am interested in that are potentially very short.
Given that the tool I use to extract the bundles is almost exclusively tested on VTK data, and does not accept TCK files, I am converting the TCK files to VTK using tckconvert ${in_fname} ${out_fname}. When I load the T1 and the resulting VTK file into 3D Slicer, streamlines land in the appropriate locations, so I assume things are OK there:
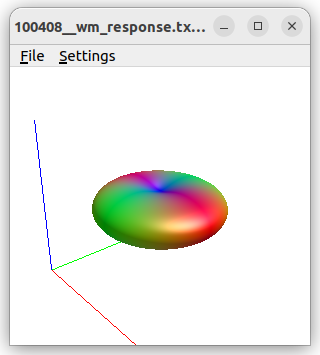
In order to do some sanity check I have turned to HCP data; I have selected the b1000 shell, computed the response function using the Tournier method,
which leads to seemingly OK FOD data:
Using iFOD2 here as well, this time with
seed_count=100000
step_size=0.7
min_length=50
max_length=340
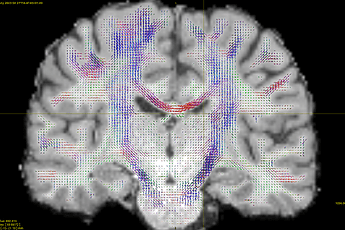
and seeding the entire WM. Note that I changed the min length here to try to see if that is a problem for the high res data/see if I get better results for HCP. Changing the cutoff down to 0.04 improves results only a tiny bit, but overall I still get poor results for the cerebellar region (below). The first thing that strikes me is that the MCP is unable to properly contour the pons:
Am I missing something in here?
Thank you.