Hi all,
I am doing the data preprocessing, and I want to create a mask for further steps. However, the created mask after bias correction does not look descent. Some non-brain areas are included in the mask. Previous steps before creating mask are:
dwidenoise dwi.mif dwi_den.mif –noise noise.mif
mrdegibbs dwi_den.mif dwi_den_unr.mif –axes 0,1
dwipreproc dwi_den_unr.mif dwi_den_unr_preproc.mif –pe_dir AP –rpe_pair –se_epi b0_pair.mif –eddy_options “ --slm=linear”
dwibiascorrect -ants dwi_den_unr_preproc.mif dwi_den_unr_preproc_unbiased.mif -bias bias.mif
dwi2mask dwi_den_unr_preproc_unbiased.mif mask_den_unr_preproc_unb.mif
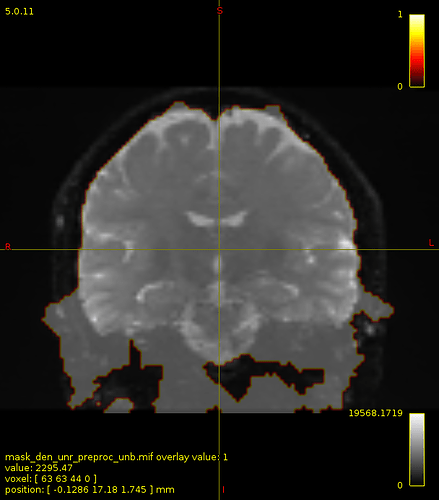
Here is the mask created after bias field correction :
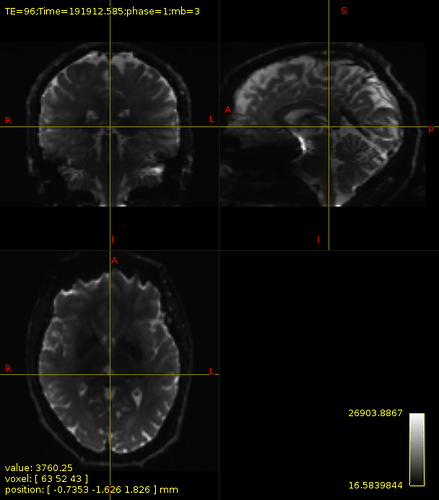
Also, I applied
dwi2mask to both unprocessed data and preproc (before biascorrection) data, as suggested by other post. The results did not differ. Creating a mask from preproc data introduces a large hole in the the midbrain.mean b0 image:
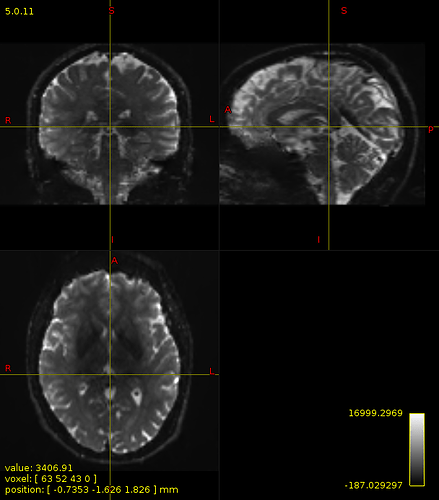
Diffusion image after biascorrection :
Except bet from fsl, is there any other way to create mask from mrtrix?
Does it mean that output of biascorrection is also invalid to be imported as the input for the next step? (e.g. remaining bias field). I mean, maybe there is a problem in the preprocessing steps which could be problematic for further analysis.
Thanks.