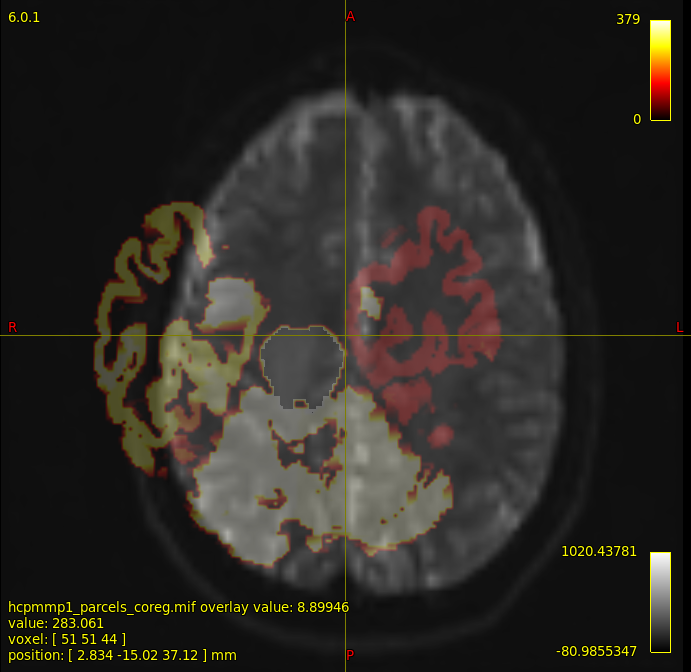
I have a problem with registration of atlas to diffusion space, i have tried doing
mri_aparc2aseg --old-ribbon --s KON11 --annot hcpmmp1 --o hcpmmp1.mgz
mrconvert –datatype uint32 hcpmmp1.mgz hcpmmp1.mif
labelconvert hcpmmp1.mif hcpmmp1_original.txt hcpmmp1_ordered.txt hcpmmp1_parcels_nocoreg.mif
mrtransform hcpmmp1_parcels_nocoreg.mif –linear diff2struct_mrtrix.txt –
and i still find improper registration
mrview mean_b0_preprocessed.mif -overlay.load hcpmmp1_parcels_coreg.mif
could some suggest me why my registration doesn’t work
Thanks
iwater
August 1, 2019, 2:28pm
2
Hi Vasudev,
Did you already align the dwi with the t1? Because the image you show looks like a problem I had some time ago. You might want to have a look at that topic, maybe it helps.
Dear MRtrixers,
After I preprocessed all my subjects. I’m trying to align my dwi data to the t1.
I did that following the steps from the tutorial but it’s not giving me any good results. I’m hoping I did something very simple wrong and I’ve been trying to figure out the problem, but so far no success.
What I did:
flirt -in mean_b0.nii.gz -ref T1.nii.gz -interp nearestneighbour -dof 6 -omat diff2struct_fsl.mat
transformconvert diff2struct_fsl.mat mean_b0.nii.gz T1.nii.gz flirt_import diff…
When that registration went well, I had no trouble with registration of the atlas to the diffusion space. Hope it helps!
Ida