Hello!
After acquiring diffusion weighted MRI data, and converting the DICOM to .nii.gz files, we found that the axes in our volumes are wrong: when displaying the .nii.gz volumes, the y- and z-axis are exchanged and the z-axis has a flipped sign.  We are now trying to reorient the images while maintaining the correct b-vectors. Our code is the following:
We are now trying to reorient the images while maintaining the correct b-vectors. Our code is the following:
mrconvert -json_import dwidata.json -fslgrad dwidata.bvec dwidata.bval dwidata.nii.gz dwidata.mif
mrconvert -axes 0,2,1,3 dwidata.mif dwidata_axisswap.mif
mrtransform -flip 2 dwidata_axisswap.mif dwidata_axisswap_flip.mif
mrconvert -strides 1,2,3,4 dwidata_axisswap_flip.mif dwidata_axisswap_flip_strides.mif
If we now extract the bval and bvec and re-convert the data to nifti
mrconvert -export_grad_fsl reorient.bvec reorient.bval dwidata_axisswap_flip_strides.mif dwidata_axisswap_flip_strides.nii.gz
and run dtifit on the data
dtifit -k dwidata_axisswap_flip_strides.nii.gz -r reorient.bvec -b reorient bval -m mask mask.nii.gz -o reorient
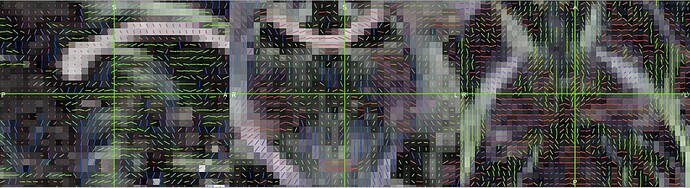
the principal diffusion direction do not seem to be correct!

(this is the V1-file from the dtifit overlaid on the FA-volume)
So it seems that the bvecs were not transformed correspondingly with the image axes…
We are happy about any suggestions, ideas, comments!