Hi,
I have a nifti file containing ROIs (labeled lesions). The labels range from 1 to 28. When I add the nifti file as an overlay on a structural MR image in mrview, not all ROI are shown. In particular the ROI with label 2 is not visible, all other ROI are visible, but not the label 2. It is not due to scaling or thresholds in the overlay panel, and also not the ‘interpolate’ option.
The only thing that works to get the ROI with label ‘2’ visible is to choose the colour map ‘Complex’. Any idea how this is possible and what I can do to make sure that the ROI is also visible with regular colour maps like ‘Jet’?
These are the ‘mrinfo’ details of the nifti file with ROIs:
Dimensions: 128 x 384 x 384
Voxel size: 1.2 x 0.677083 x 0.677083
Data strides: [ -1 2 3 ]
Format: NIfTI-1.1 (GZip compressed)
Data type: unsigned 16 bit integer (little endian)
Intensity scaling: offset = 0, multiplier = 1
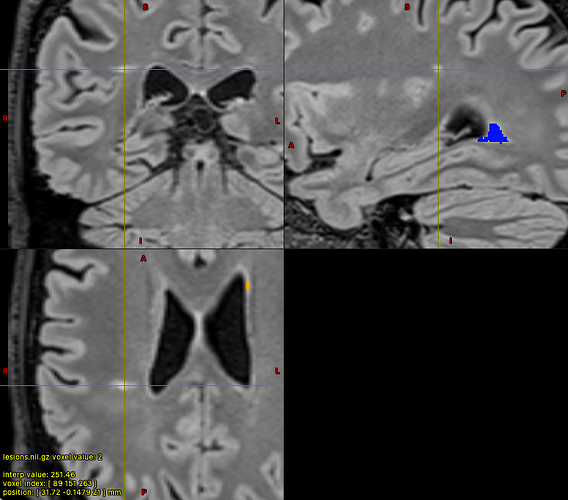
With colour map ‘Jet’
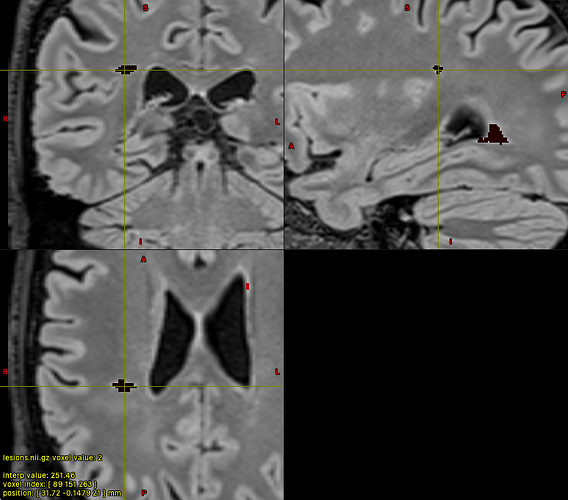
With colour map ‘Complex’
You are probably using an M1/M2 Mac and are affected by this issue: mrview: ROI edits deleted after two elements drawn · Issue #2319 · MRtrix3/mrtrix3 · GitHub
Unfortunately, there is no known workaround except for switching to an Intel Mac or Windows/Linux.
Hi Ben,
Thanks for your reply. Actually no, I still have an Intel chip.
I’m using the most recent version of mrview (3.0.4 64 bit release version, built Dec 14 2022)
I was really puzzled when I saw the missing ROI in mrview, thinking I spotted a segmentation error  but in Slicer it appeared just right, weird.
but in Slicer it appeared just right, weird.
Cheers
Thibo
This is the relevant issue: mrview: uint32 and int16 overlay display issue · Issue #2399 · MRtrix3/mrtrix3 · GitHub
It affects intel macs, and is unfortunately still a mystery apparently! Maybe you can add your specs to that thread if they’re significantly different from what’s already been documented…
Good news is, since your labels only range to 28, you can convert the image to uint8, which should be unaffected by the bug.
mrconvert in.nii.gz out.nii.gz -datatype uint8
Cheers
Fiona
1 Like
Thank you Fiona, that reassures me I didn’t create a ‘bad’ nifti file, and good to know using other datatypes may help.
Kind regards,
Thibo
1 Like
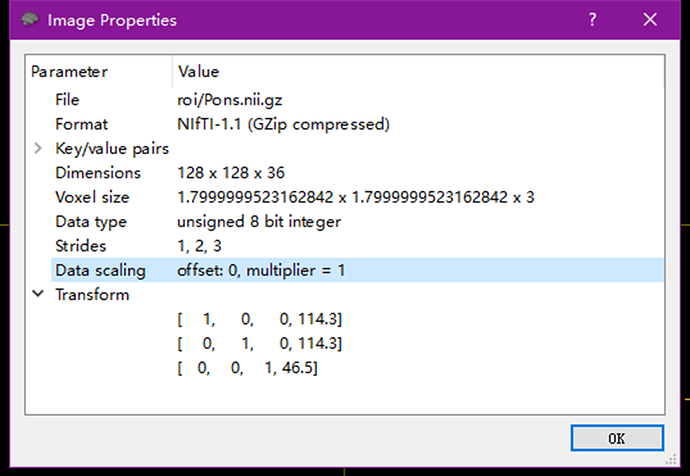
I have a similar problem, here is the ROI I generated with VTK, and I saved every spacing and origin from the original data. And the datatype is UINT8, too. In Windows10, Intel chip. But still cannot display the Pons.nii.gz in ROI mode, but load it as main image can be shown, I have no idea what’s wrong.