Hi all,
I’m somewhat of a novice with fiber tracking, looking for a bit of help.
I understand that the default value for tckgen is 100 * voxel size. When investigating interhemispheric connections, this understandably leaves very little fibers in the opposite hemisphere (400 x 400 x 264 / 0.525 mm voxelsize).
Considering fiber curvature, I am unsure how high to set this value. Will increasing it beyond a reasonable amount lead to many false positives? Any guidance appreciated.
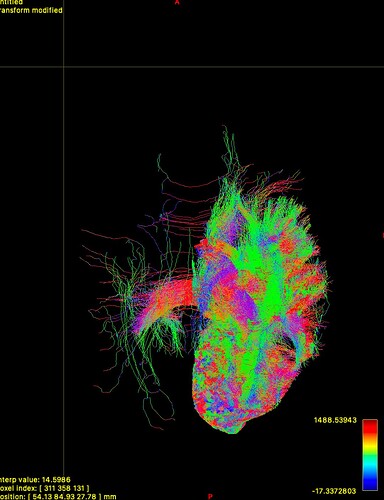
I’ve included an image as an example. Based on my reading, I would expect more fibers reaching the opposite hemisphere through the splenium here, but the default fiber length appears too short.
Thank you for reading
1 Like
I’m also curious about this question—not just for maxlength, but for other parameters as well. I’ve been investigating interhemispheric and intrahemispheric long-range connections from several lateral occipital ROIs, and I’m noticing that the projections often appear quite diffuse. For example, when looking at the endpoints of a lateral occipital seed on the lateral surface of the opposite hemisphere, the resulting connections tend to be scattered and patchy. The same holds for connections to frontal regions even in the same hemisphere.
I’ve experimented with both angle and maxlength, but neither seemed to substantially improve the tractography results. I’d be really interested to hear any recommendations for optimizing tracking of interhemispheric connections or general strategies you’ve found helpful in similar cases.