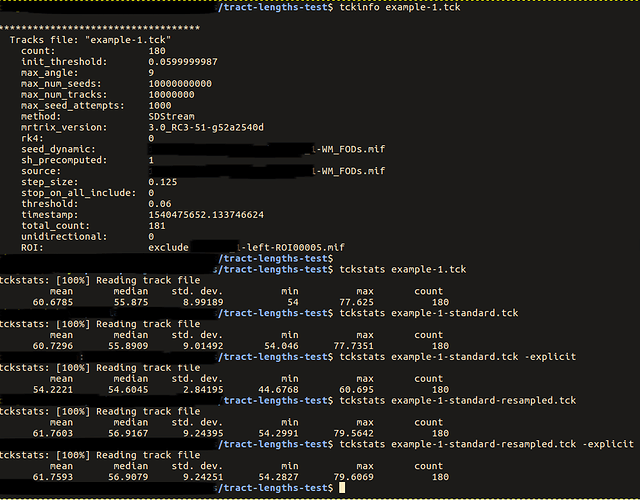
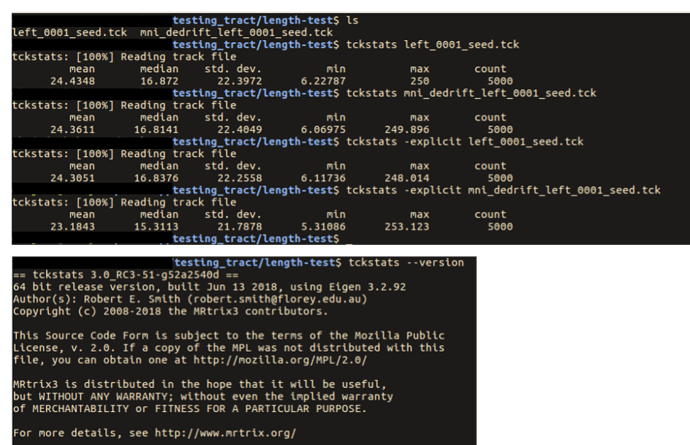
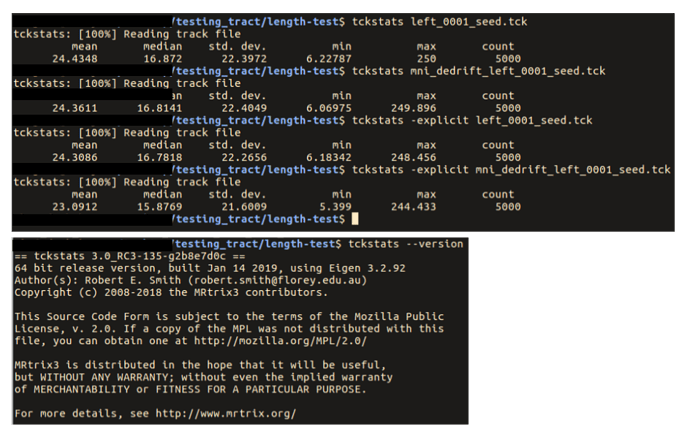
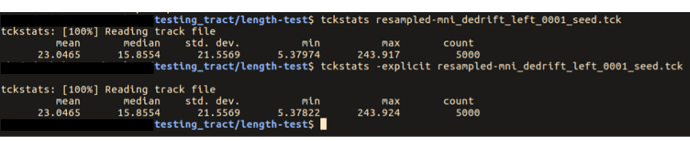
… you had mentioned that there are potential issues with tckstats with downsampled tracts …
If you’re referring to the comment here, my point there was more about the downsampling that occurs within tckgen, but it does apply to downsampling in general.
I’m not yet totally solved on how best to handle this, so I’ll describe what’s going on as best I can here in case anyone has input / writing it out in full makes my mind up.
Imagine that one possessed a method to accurately quantify the length of each streamline based on a continuous spline representation. In this case, as long as the number of discrete sampled vertices along the spline is “sufficient” (e.g. Nyquist criterion), the quantification of length should be relatively stable regardless of the actual number of vertices.
However when one calculates length based on a discrete vertex representation, whether summing the distances between vertices or taking the product of the number of vertices and the known fixed step size, the quantified length will always be shorter than the length of the continuous spline, due to using chordal lengths of curved trajectories. The fewer vertices there are, the greater the magnitude of the under-estimate.
… but this should not affect iFOD2 default downsampled. Did I understand correctly?
The reason this is not an issue for iFOD2 (specifically under default usage) is due to the internal mathematics and mechanisms of iFOD2 and the default parameters.
With default parameters, for each “step”, iFOD2 generates one new vertex at precisely (0.5 x voxel_size) distance from the current point, as well as 2 “intermediate” vertices between the current and target vertices along an arc of fixed radius of curvature. The probabilities of candidate trajectories during probabilistic streamlines tractography are calculated based on the amplitudes of the FODs at the positions and tangents defined by these four points (current point, target vertex, and two “intermediate” points). As the streamline is propagated, streamline vertices are generated at all points (including the intermediate ones), principally to ensure that any underlying ACT 5TT image or ROIs are sampled adequately densely (don’t want streamlines to “jump over” features). However by default a downsampling factor of 3 is used, which then results in all of the “intermediate” points being discarded before writing the streamline to disk.
Because the distance between the vertices at the start & end of each arc is exactly (0.5 x voxel_size), the distance between each vertex taking into account the intermediate points is in fact going to be a bit more than (1/6 x voxel_size) for anything other than a perfectly straight trajectory. However following the default downsampling, the distance between vertices will be exactly (0.5 x voxel_size).
Compare this to an alternative example: iFOD1, where the default step size is (0.1 x voxel_size), and let’s say that we use a downsampling factor of 5. Following downsampling, the distances between vertices will be less than or equal to (0.5 x voxel_size), with the magnitude of the error dependent on the radius of curvature.
For this reason, in the next tag update I’m removing all reference to calculation of streamlines lengths based on knowledge of step size.
As a slight aside: one could approximate this spline length by upsampling to a very high density of vertices along the spline before calculating the lengths. This is in fact essentially already done with tckmap -precise, tcksift, tcksift2. An issue I have is that tckgen needs to be able to apply criteria to streamlines lengths, and ideally its behaviour should be identical to applying such length criteria in e.g. tckedit, but it may be undesirable to introduce such computational demand to tckgen. I suppose I may be able to instead perform the more expensive calculation of length only if the streamline appears to be close to one of the length thresholds?
resampled step size automatically after using tcktranform (this will avoid most issues with editing transformation tracts using a length rule)
While that could be done, resampling to a fixed step size would likely be considerably slower than tcktransform itself. I would personally prefer to support piping of streamlines data so that one could run tcktransform in.tck - | tckresample - out.tck if desired. The code update linked above will prevent MRtrix3 from calculating lengths based on step size knowledge, so this will no longer be necessary in order for MRtrix3’s calculations to be accurate; the question is whether adding documentation regarding the calculation of streamlines lengths is sufficient to prevent users from making the same mistake if they choose to perform such calculations themselves manually.
remove the step size from the header after tcktranform to ensure that users cannot use this information when calculating length.
It would certainly make sense to remove (or better, invalidate) the “output_step_size” field, which, following introduction of the downsampling capability in tckgen, has been used to indicate the expected distance between vertices in the output file, as opposed to the “step size” used by the tractography algorithm. However it would be preferable to retain the latter, as it would make sense to retain this information along with all other tractography parameters.
An issue here is that when I first implemented this, there were a lot of .tck files floating around that simply didn’t contain the “output_step_size” field; in which case I would revert to using “step_size”. This in its current form would therefore perform calculations based on the “tracking” step size if the “output” step size were omitted / invalid, conflicting with the previous paragraph.
So I agree that invalidating “output_step_size” in tcktransform would be preferable, but don’t think that “step_size” should also be removed. “output_step_size” should additionally be used as a rough guide only, rather than for calculations that require precision (I think that after the update the only place that MRtrix3 code will use it will be in the determination of bin widths in a streamline length histogram).