Hi everyone,
I recently came across the slicesdir command provided by ssl package. It facilitates visual check of multiple files simultaneously. It would be great if there was something similar for mrtrix! Just a thought 
Cheers,
Vasiliki
Hey Vasiliki,
Have you looked into the multitude of command-line options now provided with mrview? These allow a range of operations - including taking a screenshot - to be automated, for instance wrapping in a script to grab one image for each of a range of subjects.
If there’s something specific about slicesdir that can’t currently be automated with mrview, could you perhaps include a screenshot and describe the specific feature(s) that you think would be beneficial for MRtrix3?
Rob
@rsmith Thanks for pointing out the command line options of mrview. How difficult would it be to add a few options to the tractography section? Specifically, we’d like to reduce the line thickness and opacity from the command line to batch capture tractography results.
Secondly, is there some way to do the rendering without a display, i.e. on our cluster? I somehow doubt it because mrview uses OpenGL (but I wanted to ask).
Thirdly  do you take pull requests for these sorts of features?
do you take pull requests for these sorts of features?
Thanks,
-dan
It shouldn’t be too hard - just need to add the option and how to handle it at this point in the code, and either invoke the existing slots or use a similar approach in the parser.
As you guessed: no. We’ve thought about implementing some kind of headless mode, but it would still require an OpenGL 3.3 stack to be present - and it seems like a lot of work to implement…
Most definitely! We are very open to others working on the code and submitting improvements - go for it!
@jdtournier PR is here: https://github.com/MRtrix3/mrtrix3/pull/946
Works like a charm.
We’re trying to add a command line option for loading a tsf for colour and thresholding.
We can access the loaded Tractogram object, and set the intensity_scalar_filename field. Calling colour_by_scalar_file_slot however fails, we think, because the object is not selected in the GUI. We can’t figure out how to do that programmatically.
Can you please point us in the right direction? Thanks!
Just thought I’d draw you attention to this recent post - seems we already might have a solution that would work, at least on Linux systems…
Something like this might do what you’re after:
$ mrview -platform offscreen anat.mif -size 1024,600 -plane 2 -mode 4 -noannot -capture.grab -exit
Great to see people getting stuck in and getting their hands dirty! ![]()
Hard to say what might be going on without looking at the code. The simplest thing to do might be to commit your changes to your own branch and create a pull request for us to look at. You can use GitHub’s fork and pull request features for this, works a treat. Here’s a little primer for those not familiar with the process:
-
make sure
gitis set up with the right details. -
create your own fork of the MRtrix3 repo (hit the fork button at the top right)
-
clone your fork: in a local terminal, type
$ git clone git@github.com:/yourusername/mrtrix3.git mrtrix3_myfork. This’ll download your changes into themrtrix3_myforkfolder. -
make your changes to the code. You can
./configure,./build, and run the code as normal, although since it won’t be in your PATH, you’ll need to invoke the executables using their path (e.g.release/bin/mrviewif you’re in your fork folder, or~/mrtrix3_myfork/release/bin/mrviewif you’re elsewhere), or update your PATH to include the relevantrelease/binfolder. -
commit your changes:
git commit -a -m "a meaningful commit message". This just records the changes you made since the last commit. You can do this as many times as required, it’s generally a good idea to add lots of small commits rather than a single giant one… (note this assumes that you’ve only modified existing files - if you’ve added new files, you’d need togit addthem explicitly first). -
push your changes to GitHub when you want to:
git push. Your changes will remain on your own GitHub fork at this point. Again, you can do this as often as you need to: if you add more commits, you can push them to GitHub whenever you’re ready - all this does it to synchronise your local changes with what’s in your GitHub fork. -
When you’re ready, create a pull request: head over to your GitHub fork on the website (it’ll be on your own account, somewhere like https://github.com/yourusername/mrtrix3), and you should see your latest commit just above the file listing. It’ll say something like “This branch is 3 commits ahead of MRtrix3:master”, and on that line there will be a link to create a pull request. Just hit that and the next ‘create pull request’ button, add as much information as you need, and then hit the ‘create pull request’ button at the bottom.
-
We’ll be notified, and then we can comment or edit the code as required.
Look forward to seeing what you’ve cooked up!
(For good record keeping)
We got this sorted. The pull is here:
Hey Rob,
It seems like I haven’t checked this thread for a while… 

If I am honest I have not tried several options of mrview-and I probably should…
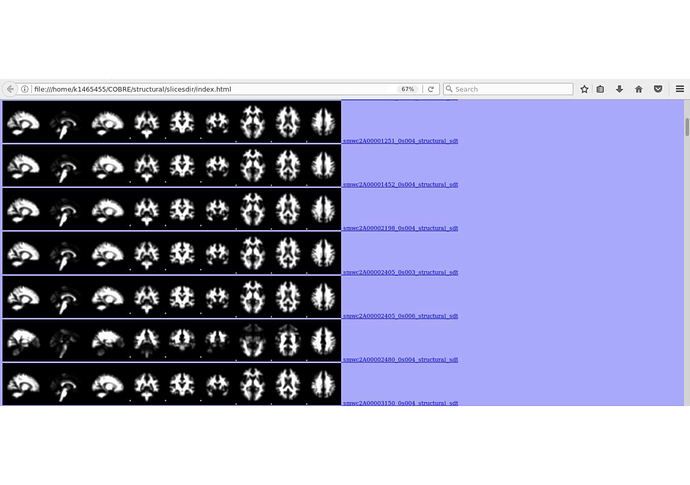
I only recommended the command slicesdir because I found it very useful for quick visual check of the whole cohort at any stage of the analysis. It essentially generates cross sections from the axial, coronal and sagittal views and generates a link that you an open in your browser. You may want to look in more detail in FSL wiki. Attached there is an example.
Cheers,
Vasiliki
Dear Vasiliki,
I also personally find view from slicesdir useful, however, in my experience, this view of image montage allow to capture only gross errors. For more detailed and reliable quality check of registration (and other processing steps) I prefer subject-wise view in fslview or mrview where also suspicious areas can be interactively viewed in more detail.
As an example, I would not know what to think of the quality of the data of the subject 2480 (frontal areas) you showed in the montage, unless I look at it in more detail in mrview.
I’m not sure that simply duplicating the operation of slicesdir would serve much purpose. It’s possibly the sort of thing that the MRtrix3 community could have an open discussion about what would be useful & widely adopted in terms of producing summary images for visual QA, and I’d be happy to provide assistance to anybody willing to take on some scripting development. However it’s also the kind of side-project that could have some major scope creep if trying to support multiple people’s preferences regarding what should be produced, so there would need to be at least some semblance of consensus.
Such a script would also benefit from me finally getting around to addressing GitHub issue #4, which would provide a binary for exporting an image slice as a .png, and thus it would be possible to generate an output like those in slicesdir without ever invoking mrview.