Dear Mrtrix experts, I folllowed the BATMAN tutorial (with few changes) to generate a whole-brain conectome based on DK atlas , which was an 84x84 matrix. And I found there was a huge number of streamlines connecting bilateral cerebellum cotex, and I don’t know what’s wrong during the analysis process.
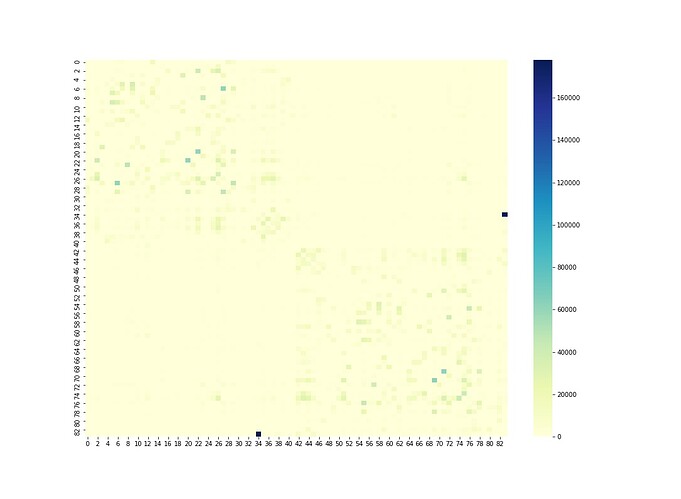
here is the connectome matrix of subject 10 (node35-84 correpond to left and right Cerebellum cortex)
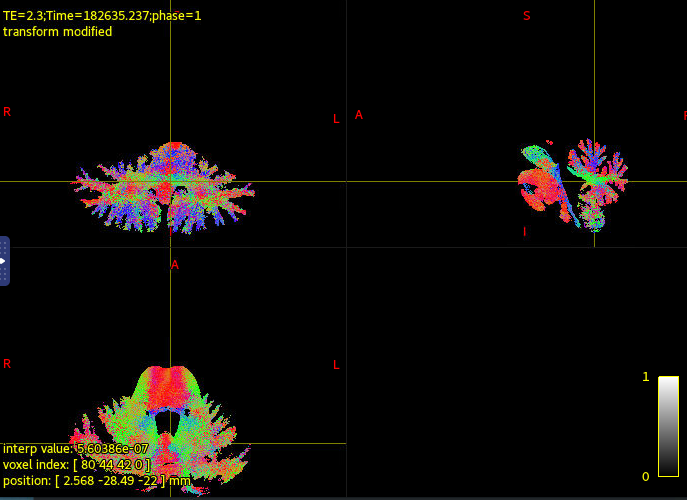
I extract the streamlines between two cerebellum using connectome2tck and here is what I am getting
Here are the related tckgen and tck2connectome commands I used:
This error is common among all subjects
I would appreciate some advice about the unexpected problems.
Best,
Kitty Zhu
Hi Kitty,
I’ve seen this in my own data in the past. There’s a good chance that this result is entirely appropriate given the data provided. If particular nodes in the parcellation are much larger in size than others, then there’s a good chance that they will have a large number of streamlines assigned to them; and if there’s two such parcels that are connected directly via a reasonably large WM pathway, then one would expect that the specific connection between them will be quite a lot more dense than other edges in the connectome.
What you probably want to be looking for is streamlines that connect the two cerebellar hemispheres, but that don’t go through those pathways that are known to connect them (i.e. MCP). For instance, there is some chance that with the combination of tissue segmentation & parcellation that you have, there are some streamlines that erroneously cross the mid-sagittal plane directly; but you would need to be confident about whether or not such connections are in fact artifactual (my own knowledge of the anatomy here is limited).
So in general, I wouldn’t be concerned about this; unless you are intending to use the density of the strongest connection in the matrix to normalise intensities across subjects (which I have seen performed elsewhere), in which case variance in this one connectivity estimate could have a large impact on the results of the entire analysis.
Rob
1 Like