Dear Stella,
your observation is probably related to the fact that there is interpolation on by default in mrview.
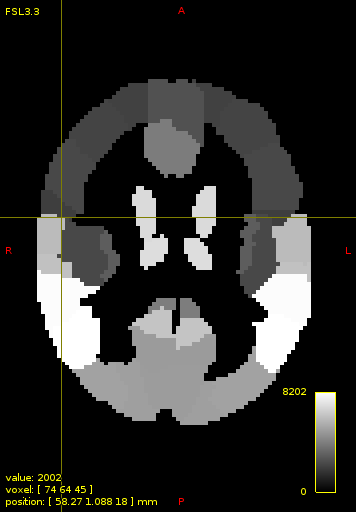
By switching interpolation off the voxels in my case contain only integer values and mrview screenshot of loaded AAL looks like this:
Antonin