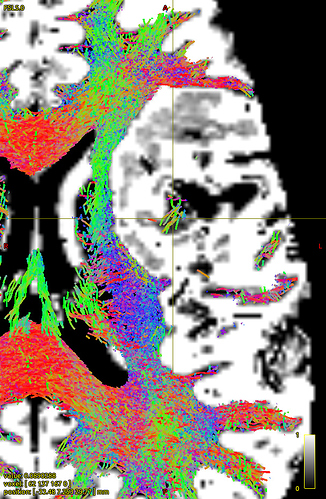
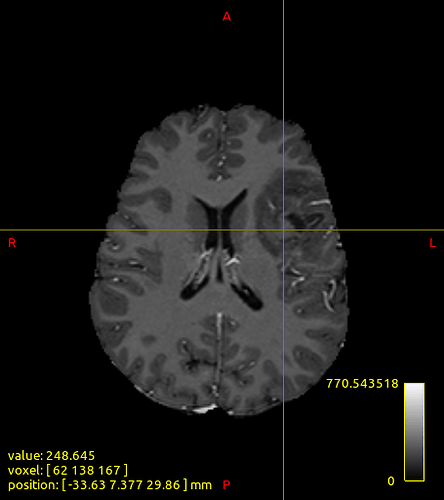
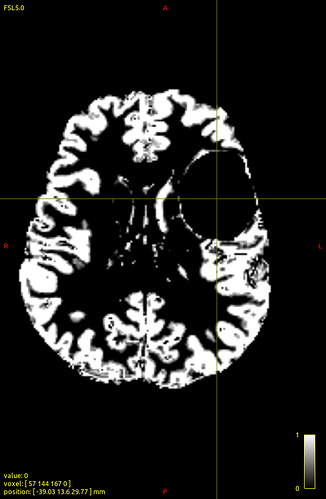
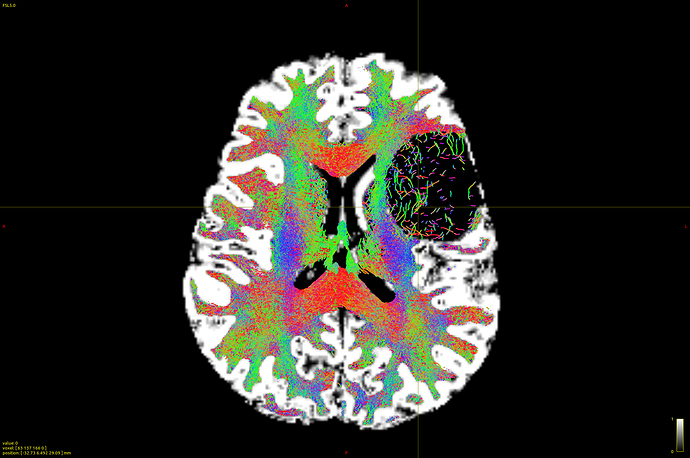
How does -seed_gmwmi take into consideration the pathological tissue volume of the 5TT image when the gmwmi image is produced from said 5TT image? Does it lower its contribution to the tract propagation from a region where voxels lie between “healthy” tissue and the lesion?
That’s … a deceptively complex question.
Upstream of -seed_gmwmi, if specifically 5tt2gmwmi is being used to produce that seed image, then the intensity of that image (which modulates the density of candidate seed points) is based primarily on this line; so if there’s an intersection between all of GM (either cortical or sub-cortical), WM, and pathological tissue, then the image could be bright and there could be a lot of seeds there (potentially an erroneously large amount, but I’d need to see evidence of such being a problem to warrant addressing).
For -seed_gmwmi itself, over and above the rejection sampling that is used to draw more or less candidate seeds in different locations, that candidate seed then has to be revised to a position that is deemed “acceptible” as a GM-WM Interface seed according to this line. Precisely how the localisation and classification of such points may behave in the presence of a delineation of pathological tissue is not something that I’ve tested.
It’s important to establish that while ACT receives as input a partial volume image, those partial volumes are not actually interpreted as “partial tissue behaviours” (as is done in this alternative). Rather, those partial volumes are simply used so that the isocontours (where the “dominant” tissue switches from one to the other) are not constrained to the voxel grid, but can approximate the hard interfaces between tissues that lead to partial volume when observed on an image grid, and can therefore be accessed very quickly using image interpolation.
It’s not clear exactly what you intended by “between” in your second question. If this is literally having healthy tissue in voxel 1, 100% pathological classification in voxel 3, and you’re asking about the behaviour purely in voxel 2, then for any sub-voxel position within voxel 2 the effect of voxel 3 due to interpolation has essentially “ended” (due to the interpretation of hard boundaries rather than partial tissue behaviours). If instead you’re asking about individual voxels where the pathological classification is not binary but the volume of that voxel is distributed “between” pathological and healthy tissue, the presence of non-zero pathological classification is only applicable in sub-voxel positions where, following interpolation, the pathological tissue is the dominant one.
If you intended something different, you’d need to be more specific; if I’m ever going to get on top of this forum, I can only get so far into the weeds.
Cheers
Rob