Hello everyone,
I am working on ex-vivo diffusion data acquired on a part of the brain centered on the basal ganglia at 11.7T.
The acquisition is as follows, multishell B1000 7 dirs B4000 29 dirs B10000 64 dirs.
I did the following pre-processing steps:
# Concatenate all the DWI images and the B0 map into a single 4D image
mrconvert 64dirsB10000/v_1.nii processed/B10000.mif -fslgrad 64dirsB10000/bvecs 64dirsB10000/bvals #OK
mrconvert 29dirsB4000/v_1.nii processed/B4000.mif -fslgrad 29dirsB4000/bvecs 29dirsB4000/bvals #OK
mrconvert 7dirsB1000/v_1.nii processed/B1000.mif -fslgrad 7dirsB1000/bvecs 7dirsB1000/bvals #OK
mrcat processed/B10000.mif processed/B4000.mif processed/B1000.mif processed/dwi.mif #OK
## Add dwidenoise :
dwidenoise -mask processed/T2star_brain_mask.mif processed/dwi.mif processed/dwi_dwidenoised.mif
# Correct for Eddys current distortions and motion
dwipreproc processed/dwi.mif processed/dwi_corrected.mif -rpe_none -pe_dir AP -eddy_options " --slm=linear --data_is_shelled"
dwipreproc processed/dwi_dwidenoised.mif processed/dwi_dwidenoised_corrected.mif -rpe_none -pe_dir AP -eddy_options " --slm=linear --data_is_shelled"
# Fit DTI model to the concatenated image using MRtrix3's dwi2tensor command
dwi2tensor processed/dwi_corrected.mif processed/tensors.mif
dwi2tensor processed/dwi_dwidenoised_corrected.mif processed/tensors_denoised.mif
# Calculate FA, MD and other scalar maps using MRtrix3's tensor2metric command
tensor2metric processed/tensors.mif -fa processed/FA.mif -type rgb processed/FAColor.mif -mask processed/T2star_brain_mask.mif -force
tensor2metric processed/tensors.mif -fa processed/fa.mif -adc processed/md.mif -vector processed/dec.mif -mask processed/brain_mask_bin.mif -force
tensor2metric processed/tensors.mif -adc processed/ADC_MD.mif -mask processed/T2star_brain_mask.mif -force
tensor2metric processed/tensors_denoised.mif -fa processed/fa_denoised.mif -adc processed/md_denoised.mif -vector processed/dec_denoised.mif -mask processed/brain_mask_bin.mif -force
dwi2response dhollander processed/dwi_dwidenoised.mif processed/response_wm.txt processed/response_gm.txt processed/response_csf.txt -mask processed/brain_mask_bin.mif -voxels processed/voxels.mif -nthreads 75 # OK WORKED yeeeeeee
dwi2fod msmt_csd processed/dwi_dwidenoised.mif processed/response_wm.txt processed/wm_FOD.mif processed/response_gm.txt processed/gm_FOD.mif processed/response_csf.txt processed/csf_FOD.mif -mask processed/brain_mask_bin.mif -nthreads 75
mrconvert -coord 3 0 processed/wm_FOD.mif - | mrcat processed/csf_FOD.mif processed/gm_FOD.mif - processed/vf.mif
mrview processed/vf.mif -odf.load_sh processed/wm_FOD.mif
mrconvert processed/T2star_brain.nii.gz processed/T2.mif
5ttgen fsl processed/T2.mif processed/5tt_nocoreg.mif -t2 processed/T2.mif -premasked #useless ... ?
mrview processed/5tt_nocoreg.mif
And from it I extracted fibers of interest using masks made by other imaging modalities
for example :
tckgen -force -seed_dynamic wm_FOD.mif -nthreads 60 -maxlength 250 -cutoff 0.1 -select 100000000 wm_FOD.mif tracks_100M.tck
tckedit -include $mask1 -include $mask2 Nicolas_Tempier/DTI/masks_for_dti_mif_registered/GPe_GPi_V4/tracks_100M_$(basename "$mask1" .mif)_$(basename "$mask2" .mif)_raw_lengths.tck -maxlength 300 -minlength 0.001 -force -nthreads 78
So my question is : in terms of pre-processing, how can I optimize/denoiee my datas please ?
I feel like it is a bit hard to find ex-vivo optimised preprocessing lecture,
I found this really good one :
Recommendations and guidelines from the ISMRM Diffusion Study Group for preclinical diffusion MRI: Part 2 – Ex vivo imaging
But it does not really goes into preprocessing specificities
I’ve read the forum posts too :
This post was really useful for me :
But they are getting old, new versions and algorithms arrived in MRtrix3, what would you recommend ?
Thank you in advance for your feedback,
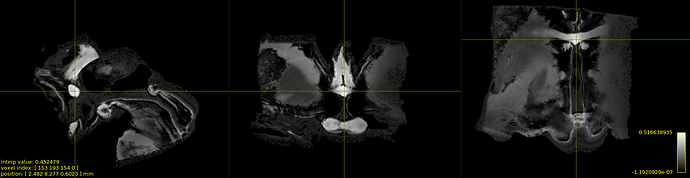
if needed, here are some results I got with this setting:
WMFOD :
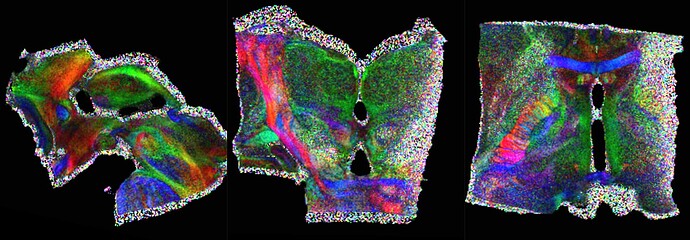
Coloured FA :
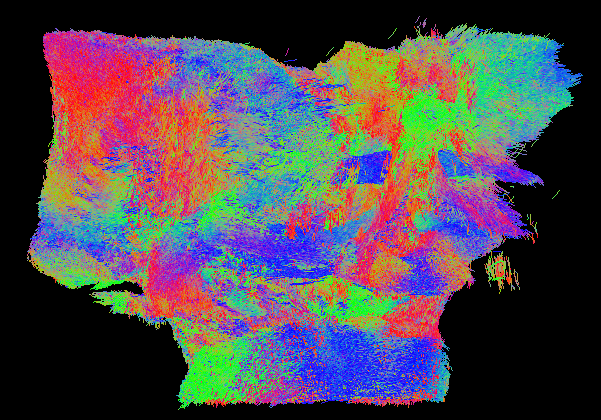
Full specimen 200k tracks :
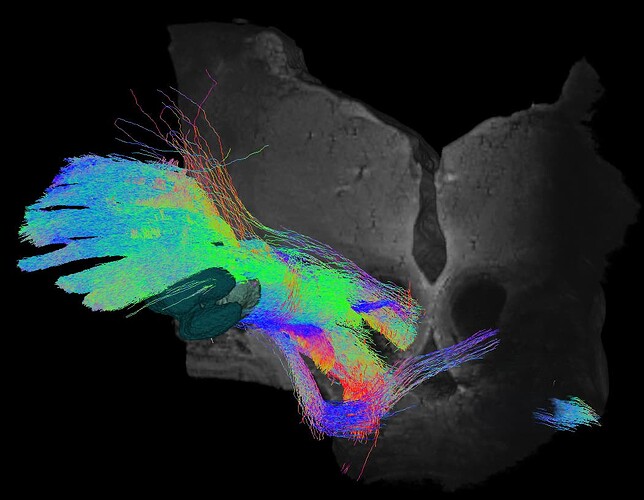
Mask related tracks using eGP iGP STN masks ans 100M tracks :