Hi Mrtrix community,
We tried to register the whole-brain tractogram to MNI space, but some problems appeared.
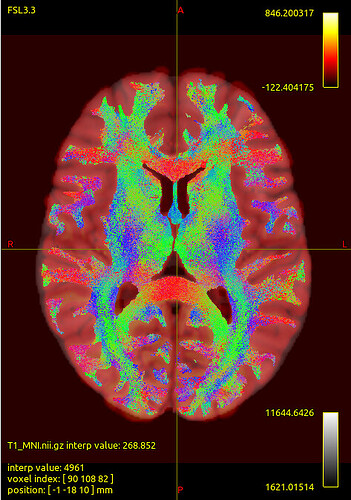
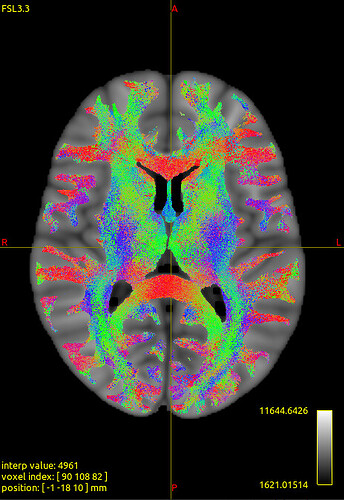
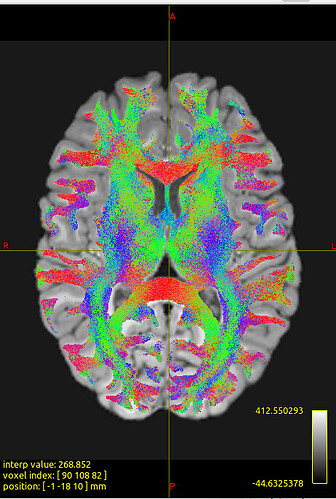
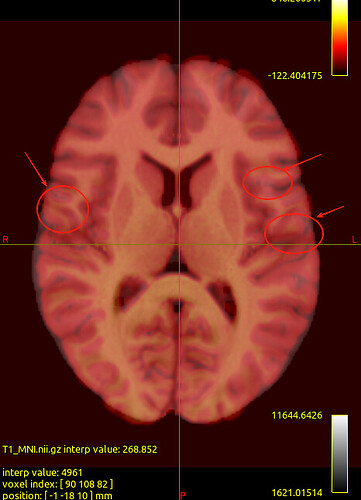
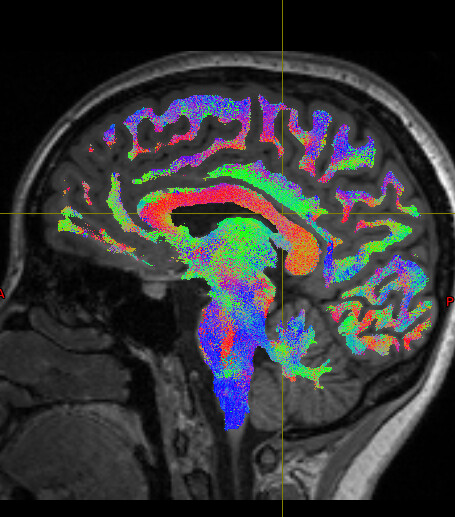
We noticed that the tck files already warped to MNI space are misaligned with the standard template in some sulcal and gyral regions. However, we also checked the registration between the T1 image and MNI space — even the T1 image warped to MNI space fails to align with the sulci and gyri of the MNI template. Nevertheless, the tck files in standard space show better alignment with the T1 image in MNI space. Can this be considered a good warping result? Figures are as follows.
My script for registration and warping
DWI to T1
warpinit Diff2MNI/unskull_T1w.nii.gz Diff2MNI/flirt-.mif
mrtransform Diff2MNI/flirt-.mif -linear xmfs/T_DWItoT1.txt Diff2MNI/flirt2tck.mif -inverse
tcktransform Tracks_1m.tck Diff2MNI/flirt2tck.mif Diff2MNI/Tracks_T1.tck -force
T1 to MNI
According to https://community.mrtrix.org/t/registration-using-transformations-generated-from-other-packages/2259
antsRegistrationSyN.sh -d 3 -f ${FSLDIR}/data/standard/MNI152_T1_1mm_brain.nii.gz
-m Diff2MNI/unskull_T1w.nii.gz -o Diff2MNI/ants/T12Standard -t ‘s’ -n 24
antsApplyTransforms -d 3 -i Diff2MNI/unskull_T1w.nii.gz -r ${FSLDIR}/data/standard/MNI152_T1_1mm_brain.nii.gz
-o Diff2MNI/T1_MNI.nii.gz
-n BSpline
-t Diff2MNI/ants/T12Standard0GenericAffine.mat
-t Diff2MNI/ants/T12Standard1Warp.nii.gz
warpinit ${FSLDIR}/data/standard/MNI152_T1_1mm_brain.nii.gz Diff2MNI/inv_identity_warp.nii -force
for i in {0..2}; do
antsApplyTransforms -d 3 -e 0 -i Diff2MNI/inv_identity_warp${i}.nii
-o Diff2MNI/inv_mrtrix_warp${i}.nii
-r Diff2MNI/unskull_T1w.nii.gz
-t [Diff2MNI/ants/T12Standard0GenericAffine.mat, 1]
-t Diff2MNI/ants/T12Standard1InverseWarp.nii.gz --default-value 2147483647
done
warpcorrect Diff2MNI/inv_mrtrix_warp.nii Diff2MNI/inv_mrtrix_warp_corrected.mif -marker 2147483647 -force
tcktransform Diff2MNI/Tracks_T1.tck Diff2MNI/inv_mrtrix_warp_corrected.mif Diff2MNI/Tracks_MNI.tck -force
Many thanks!