Dear all,
We have some doubts about normalization of tck file.
we have got the tck file in the native space in line with B0 image. we also have the nonlinear registration file i.e. warp image, from the registration between B0 and MNI template with fsl. Next, we want to check the tracts in the MNI space, but meet the problem of using tcktransform. Could someone share the experience of tract registration, or what transform file should be used in the command “tcktransform in transform out”?
Many thanks!
tcktransform currently requires warps in the MRtrix3 format. See Registration: using transformations generated from other packages for how to generate and apply these and optionally here for a full worked example of tcktransform.
Hi @maxpietsch
Thanks for your reply.
I tried some code according to your reply. But they didn’t work.
I have b0 image (nodif.nii.gz), template image (MNI152_T1_1mm_brain.nii.gz), the tractography result (MNI152_T1_1mm_brain.nii.gz) and warp image got from FSL(MNI_2nodif_warp.nii.gz, nodif_2MNI_warp.nii.gz)
I want to transform the tck file to MNI space.
I tried three methods.
First, I used transformconvert and warpconvert to convert nodif_2MNI_warp.nii.gz to a warp format usable in MRtrix3, then used it to transform tck file to MNI space.
But, transformconvert allows to convert linear transformation matrices and warpconvert requires the warpfull file is 5D format. I don’t know to how to convert the warp I have got from FSL. Is there any other command available?
Second, I used the mrregister to get the warp then transform tck file to MNI space:
mrregister nodif.nii.gz MNI152_T1_1mm_brain.nii.gz -nl_warp_full warp_full.mif
warpconvert warp_full.mif warpfull2deformation w_t2s.mif -from 2 -template nodif.nii.gz
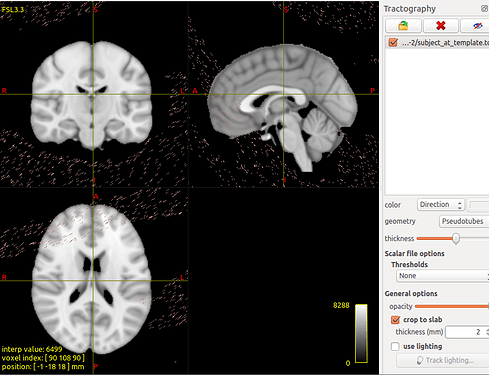
tcktransform SD_Stream_SIFT.tck w_t2s.mif subject_at_template.tck
But the result not correct.
The transformed tracts on the template:
Third, I want to try these code. But I only tried the command warpinit. Because I didn’t install ANTs and have never used it.
warpinit MNI152_T1_1mm_brain.nii.gz.nii.gz inv_identity_warp.nii.gz
inv_identity_warp.nii.gz seems to be right.
If I don’t use ANTs, how can I do that?
Stay healthy and stay happy!
Best regards,
Wu Xiao
Hi Wu,
warpconvertrequires thewarpfullfile is 5D format. I don’t know to how to convert the warp I have got from FSL. Is there any other command available?
Only the warpconvert conversion operations that start with “warpfull” necessitate 5D full warp file format. Given that format contains information over and above that of a 4D warp field, there is no way to convert from the latter to the former. warpconvert can however convert between “displacement” and “deformation” fields, which are both 4D; the difference between these two is explained in the warpconvert help page.
Second, I used the
mrregisterto get the warp then transform tck file to MNI space:
$ mrregister nodif.nii.gz MNI152_T1_1mm_brain.nii.gz -nl_warp_full warp_full.mif
This will not produce a good result, as mrregister currently only supports the squared-difference metric, and therefore cannot be used to register images of different modalities that contain different contrasts.
Third, I want to try these code. But I only tried the command warpinit. Because I didn’t install ANTs and have never used it. … If I don’t use ANTs, how can I do that?
While it may not be possible to follow the instructions from that wiki page command by command if using a registration software other than ANTs, the same logic may still apply depending on the content of the warp file that you are using, which in turn depends on the software and how it was used.
The key for you is image MNI_2nodif_warp.nii.gz. As described in the wiki post, warping streamlines requires the inverse transformation: streamline vertices are “pushed” to their new locations, as opposed to image transformation, where data for each target image voxel are “pulled” from the image being transformed. But the fact that you are provided with both “MNI_2nodif_warp.nii.gz” and “nodif_2MNI_warp.nii.gz” means that this inversion has already taken place (whether implicitly during registration, or explicitly calculated afterwards for you).
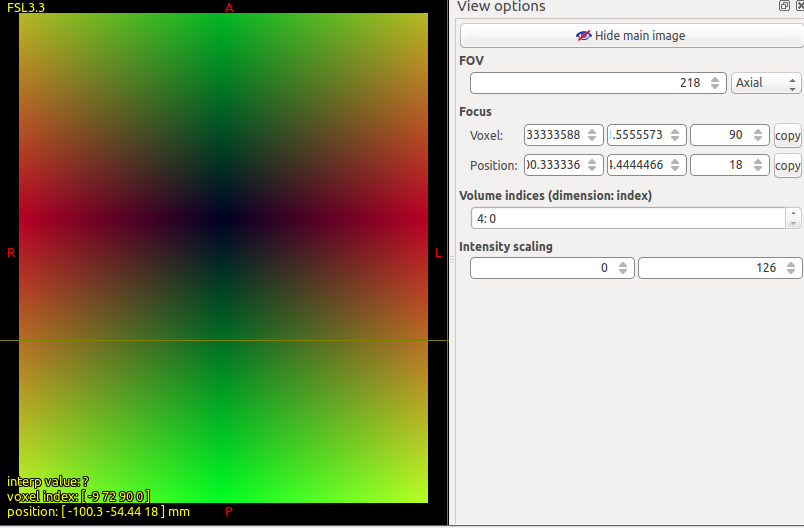
The question then is, in what format does this image provide its non-linear transformation? Simply running commands that produce the incorrect result may not be terribly useful here. It would be more direct to instead inspect the raw values that are contained within the image, to determine whether it is a displacement or deformation field, and therefore whether or not it requires conversion in order for tcktransform to interpret the data correctly. I don’t know whether it’s possible for displacement fields to be defined w.r.t. image axis spacing and/or orientations… but at least using the right input image, and the right interpretation of displacement vs. deformation, should get you close to the correct result.
Cheers
Rob
To summarise, these files are in FSL format, instead of converting them yourself, apply the correct warp to the files produced by warpinit as described in Registration: using transformations generated from other packages but using applywarp instead of antsApplyTransforms. See for instance here.
Hi,
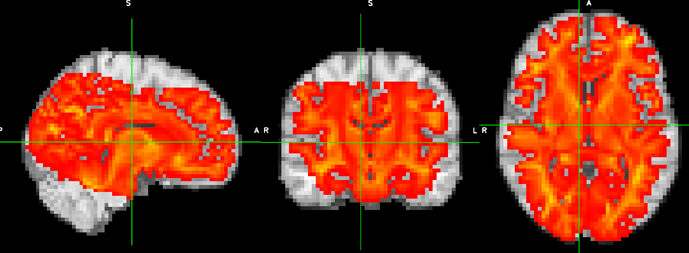
I was dealing with the similar problem: to transform the *.tck in native DWI space to MNI space using the FSL generated warps image between DWI and MNI, the results seems good:
background is the MNI_3mm_brain.nii, and the red color is the transformed tck.nii.
here are the commands I used:
#generate the MNI2DWI transformation warp by mrtrix (combined with the fsl warps: warps_MNI2dwi.nii.gz)
warpinit MNI152_T1_3mm_brain.nii.gz inv_identity_warp_no.nii
applywarp --ref=b0_brain.nii.gz --in=inv_identity_warp_no.nii --warp=warps_MNI2dwi.nii.gz --out=mrtrix_warp_MNI2dwi.nii.gz
#check the warping results by applying it to MNI_3mm_brain
mrtransform MNI152_T1_3mm_brain.nii.gz -warp mrtrix_warp_MNI2dwi.nii.gz MNI2dwi.nii.gz
#tcktransform
tcktransform ACT_WM.tck mrtrix_warp_MNI2dwi.nii.gz tck2MNI.tck -force
tckmap tck2MNI.tck tck2MNI.nii -template MNI152_T1_3mm_brain.nii.gz -force
Thanks, @LiuYuchen! If you want, I think it’d be great if you could add your minimal working example (and optionally also the command that generate the warp) as a section ## Warping tracks files to MNI space using FLS warps or similar to the end of the wiki post Registration: using transformations generated from other packages.
Glad to share my little experience ![]()
Do I need to create a new topic?
Great! In the wiki category, all posts are encouraged to be edited by anyone who can contribute something to that topic. Donald described it here.
So, to edit that post, just click the green icon
Got it 
@LiuYuchen and anyone reading this: You might want to use warpcorrect on the output of applywarp as the output of applywarp can contain 0,0,0 at locations outside the FOV of the source image. Since the image is subsequently interpreted as a deformation field, you can end up pulling image intensity values from that location or get spurious streamline vertices at locations that the original image did not cover.